VFB hosts a range of datatypes, including electron microscopy (EM) and light microscopy (LM) datasets with images annotated using the same cell type ontology terms and aligned to common standard templates. We also host single cell RNA sequencing (scRNAseq) gene expression data.
This is the multi-page printable view of this section. Click here to print.
Data on VFB
- 1: Electron Microscopy Data
- 2: Light Microscopy Data
- 2.1: Janelia FlyLight
- 3: Single Cell RNA Sequencing Data
- 4: Templates
1 - Electron Microscopy Data
Virtual Fly Brain brings together data from multiple electron microscopy (EM) resources, providing access to high-resolution neuroanatomical datasets. These datasets include complete EM volumes of Drosophila brains and ventral nerve cords, neuron reconstructions, connectivity information and neurotransmitter predictions. Data can be visualised using the web browser or accessed programmatically via APIs.
Comparison Table of Integrated Datasets
The table below summarises EM datasets that have been integrated into VFB, including the portion of the organism covered (Anatomy), the resource(s) where the original data can be found, The level of reconstruction (sparse or dense) and the original publication for the dataset.
| Dataset | VFB symbol | Anatomy | Reconstruction | Resource(s) | Original Publication |
|---|---|---|---|---|---|
| BANC | BANC | Full CNS (adult female) | Dense | Codex | Bates et al. (2025) |
| Optic-lobe | ol | Optic lobe (adult male) | Dense | NeuPrint; Codex | Nern et al. (2025) |
| FAFB (FlyWire) | fw | Full brain (female larva) | Dense | Codex | Dorkenwald et al. (2024); Schlegel et al. (2024) |
| MANC | mv | Full VNC (adult male) | Dense | NeuPrint; Codex | Takemura et al. (2024) |
| FANC | N/A | Full VNS (adult female) | Sparse | CATMAID | Phelps et al. (2021) |
| Hemibrain | hb | Partial brain (adult female) | Dense | NeuPrint | Scheffer et al. (2020) |
| FAFB (CATMAID) | fafb | Full brain (adult female) | Sparse | CATMAID | Zheng et al. (2018) |
| l1em | l1em | Full CNS (female larva) | Sparse | CATMAID | Ohyama et al. (2015) |
Datasets Hosted by VFB
Virtual Fly Brain (VFB) hosts several CATMAID instances for exploring connectomic reconstruction data. These datasets provide access to neuroanatomical data from various Drosophila electron microscopy projects. Full details are available here.
2 - Light Microscopy Data
Virtual Fly Brain brings together data from multiple Light Microscopy (LM) resources, providing access to visualisations of driver (including split) expression patterns.
2.1 - Janelia FlyLight
Introduction
FlyLight makes its primary data available via VFB as well as through dedicated websites for expression patterns of Generation 1 GAL4 and LexA lines and for split-GAL4 lines. Generation 1, split hemidriver, and stable split combination lines are available primarily from the Bloomington Stock Center. Stable split lines can also be requested via the split-GAL4 website.
FlyLight Data and Resources
FlyLight data is integrated and indexed on VFB and direct linkouts are provided from the Term Info tabs to the original images. VFB hosts the original larval images here. External links to the original collections of adult images are provided below:
Generation 1 GAL4 expression patterns
Generation 1 MCFO (single neuron) patterns
Split-GAL4 Expression Patterns
Light and EM neurons can also be searched for on NeuronBridge
Also see FlyLight Imaging Tile Documentation.
3 - Single Cell RNA Sequencing Data
Virtual Fly Brain incorporates scRNAseq data from multiple studies, including the Fly Cell Atlas, Aging Fly Cell Atlas and other datasets that identify nervous system cell types - full list here. These datasets can be found by searching for ‘scRNAseq’ and filtering to ‘Dataset’.
Clusters can be found for particular cell types via the Single cell transcriptomics data for.. query on the cell type Term Info pane.

Genes for a particular cluster can be filtered by function and sorted by expression level and extent (proportion of cells in cluster expressing the gene). As with other VFB search results, these can be exported as a csv.
Note that we currently only include genes that have extent > 0.2 in a cluster.
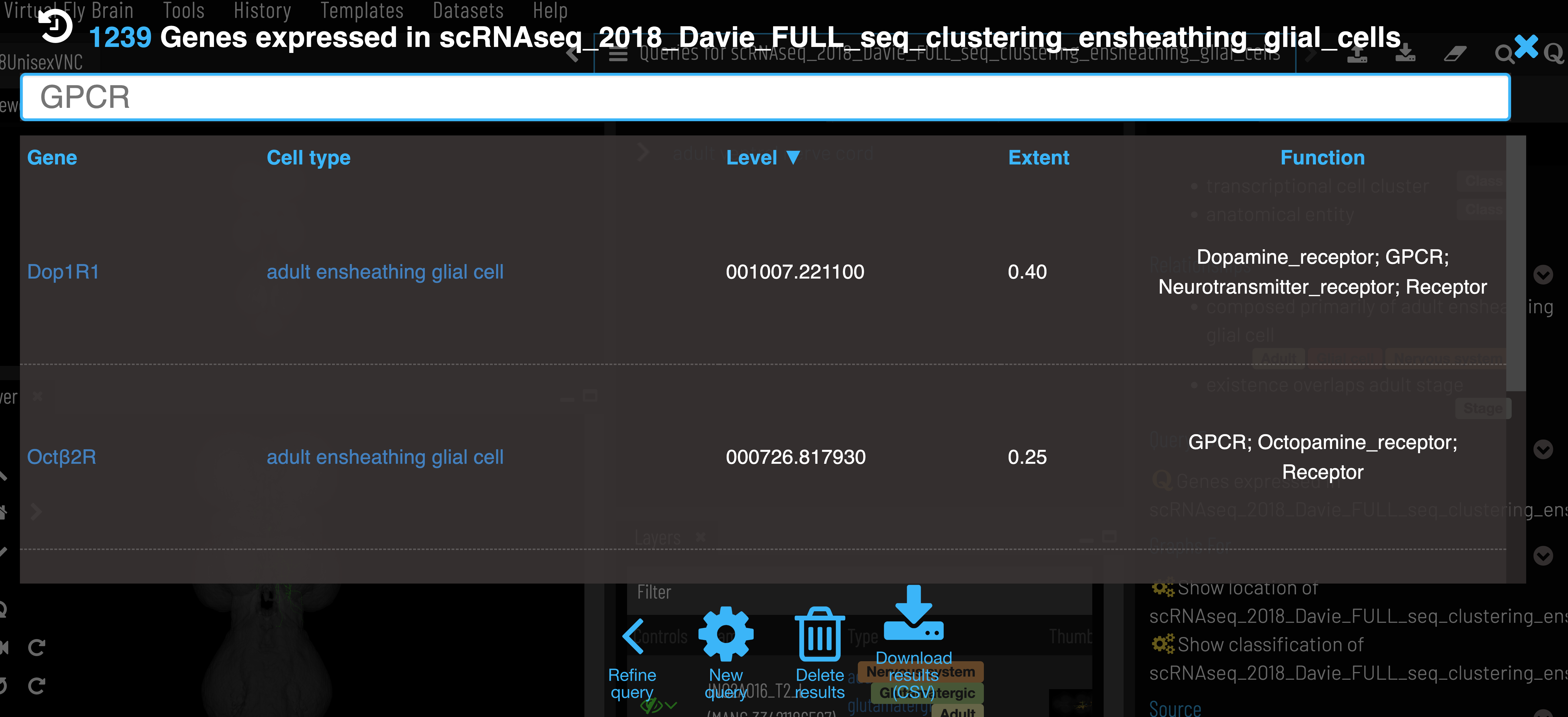
Data can also be retrieved using VFB_connect.
We pull scRNAseq data from FlyBase, which takes it from the Single Cell Expression Atlas.
4 - Templates
Canonical templates not only allow for spatial alignment of image data, but also are often painted to make a reference atlas of anatomical regions. Many central nervous system (CNS) templates exist for Drosophila, below we provide a summary of those used in VFB. Templates can be downloaded from their Term Info pages.
Central Nervous System from Janelia Research Campus JRC2018
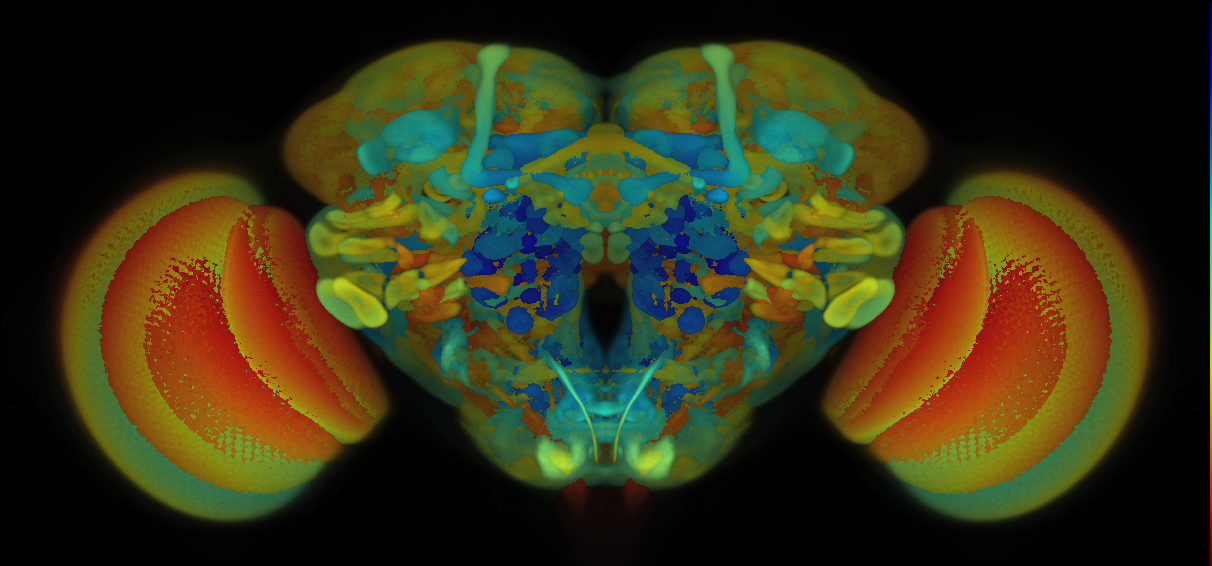
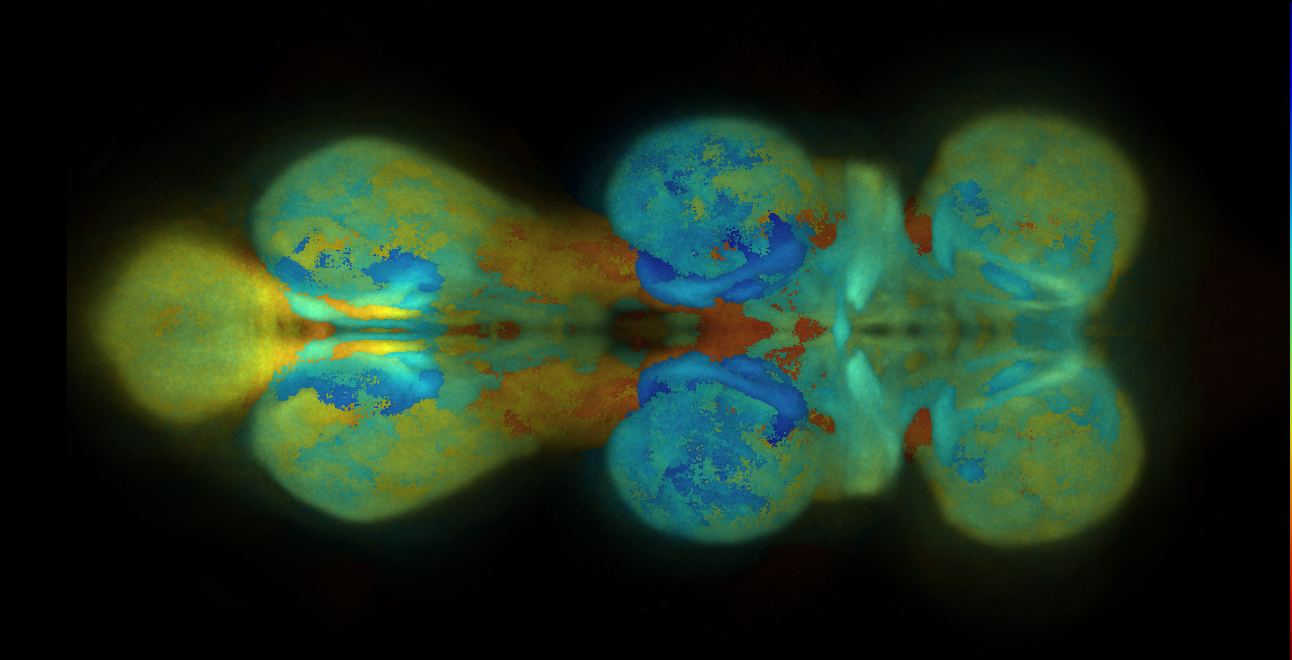
VFB displays data aligned to the JRC 2018 unisex brain template or JRC 2018 unisex Ventral Nerve Cord (VNC), as appropriate. See Bogovic et al., “An unbiased template of the Drosophila brain and ventral nerve cord” for details of template generation.
The original files are available from Janelia: JRC 2018 Brain templates.
Adult Brain from Janelia Research Campus/VFB (JFRC2010/JFRC2)
Superceeded by JRC2018
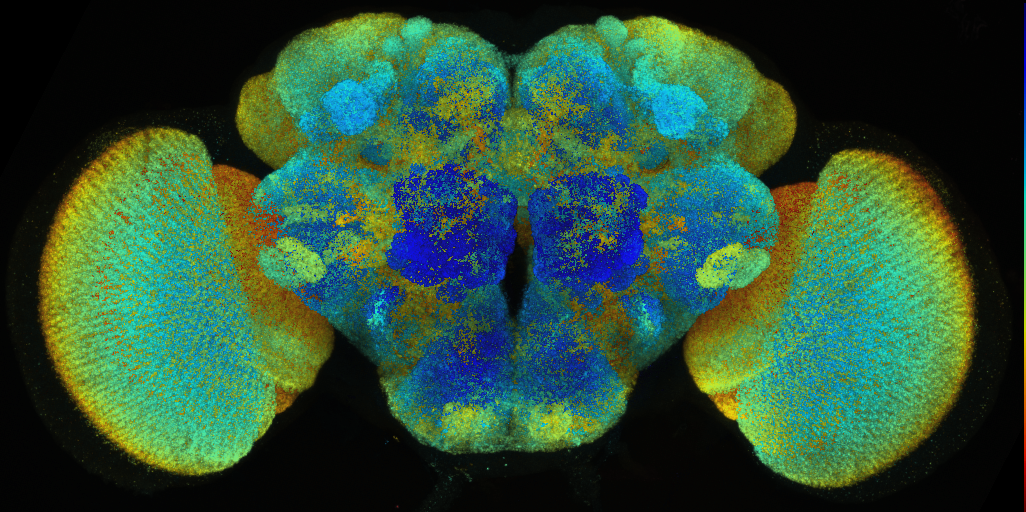
Template brain created by Arnim Jenett (Janelia Research Campus), Kazunori Shinomiya and Kei Ito (Tokyo University) from a staining with the neuropil marker nc82.
The voxel size is 0.62x0.62x0.62 microns.
The files are available here.