This is the multi-page printable view of this section. Click here to print.
Website Tutorials
1 - Similarity Score Queries Guide
Scores Availability
The similarity scores are calculated using NBLAST or other third-party scores (e.g. NeuronBridge). Only queries with above-threshold scores will appear in the ‘Find similar…’ menu.
Step 1: Select a Neuron or Expression Pattern
Navigate to the VFB browser and select a neuron or expression pattern of interest so it’s information is shown in the Term Info panel.
Step 2: Open the ‘Find similar…’ Menu under the ‘Query For’ section
Under the ‘Query For’ section, locate the ‘Find similar…’ expandable menu. This menu will only appear if an above-threshold score is available for the selected neuron or expression pattern.
Step 3: Run a Similarity Score Query
Inside the ‘Find similar…’ menu, you will find queries to find morphologically similar neurons or expression patterns. Select a query to run it. The results will display neurons or expression patterns that are similar to your selected neuron or expression pattern.

Note: We recalculate these scores shortly after new data is added, so new matches may appear if you check back a few months later.
2 - Bulk Image Download
Using the Bulk Image Download Tool
Follow these steps to download multiple files from VirtualFlyBrain using the bulk download tool:
Step 1: Open the Images
Navigate to the VFB Browser and open all the images you are interested in downloading. You can do this by searching for specific images and opening their respective pages.
Step 2: Open the Bulk Download Tool
Once you have all the images open, locate the bulk download icon in the top right. Click on this icon to open the bulk download tool.
Step 3: Select the Data Type
In the bulk download tool, select the type of data you want to download. The options are OBJ, SWC, NRRD, and References.
Step 4: Select the Variables
If applicable, select specific variables related to the data type you have chosen.
Step 5: Download the Data
After making your selections, click the ‘Download’ button to start the download. The data will be downloaded in a zip file named “VFB Files.zip”.
Step 6: Unzip the File
Once the download is complete, locate the zip file in your downloads folder and unzip it to access the data.
Step 7: Open the Data
Finally, open the data in your chosen software. For example, if you chose OBJ, you could open the data in a 3D modeling program.
Note: If something goes wrong during the download, an error message will be displayed and you will be prompted to try again. If no entries are found for the selected types and variables, a message will be displayed to inform you.
3 - Browsing single cell RNAseq data
Finding transcriptomics data on VFB
Finding cell type clusters
Cell types with available scRNAseq data can be identified using the ‘Has scRNAseq data filter’ when searching.
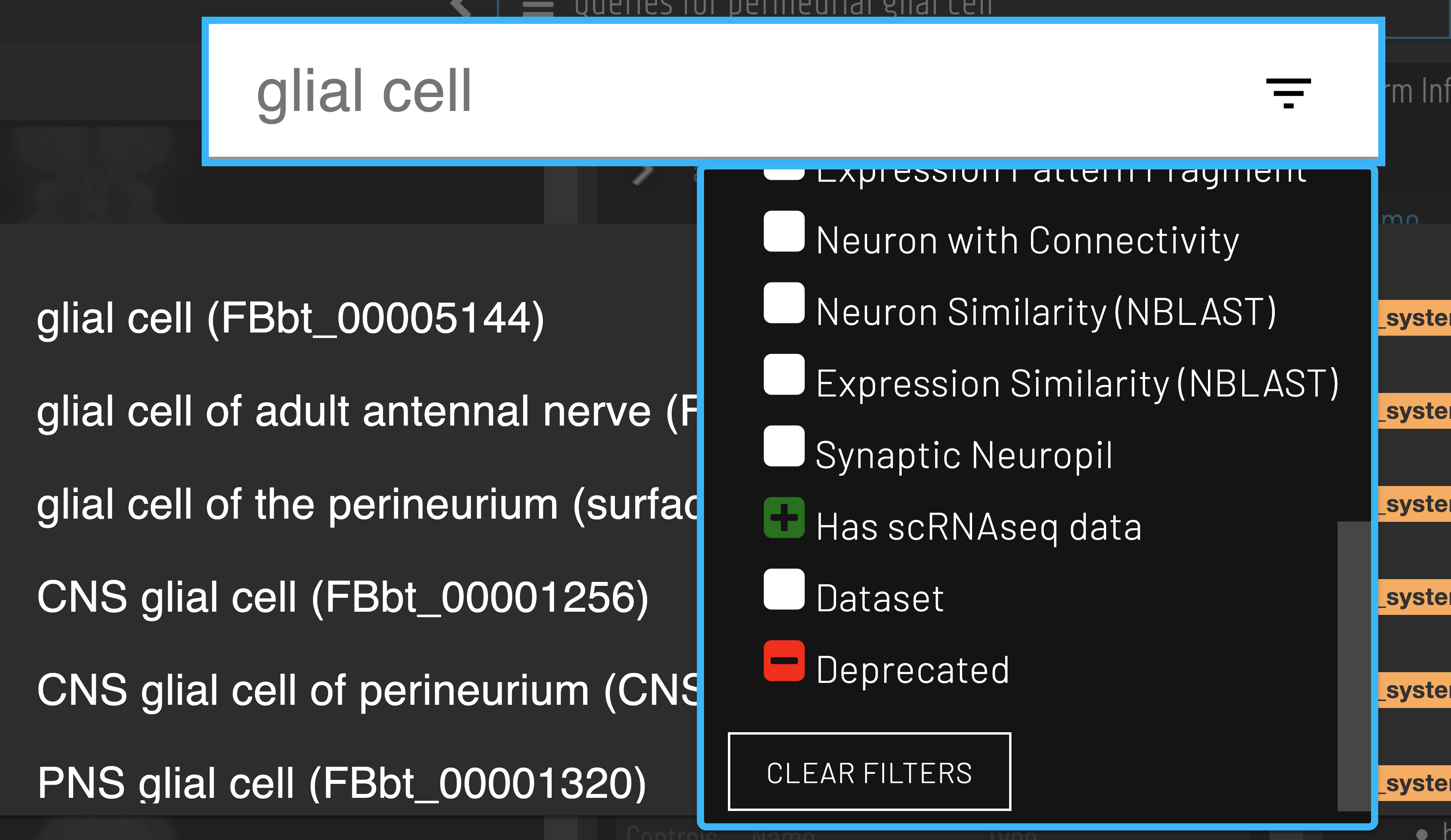
Clusters for a cell type of interest can be found via the ‘Single cell transcriptomics data for…’ query.

Gene expression data
After selecting a cluster, gene expression can be retrieved via the ‘Genes expressed in…’ query.
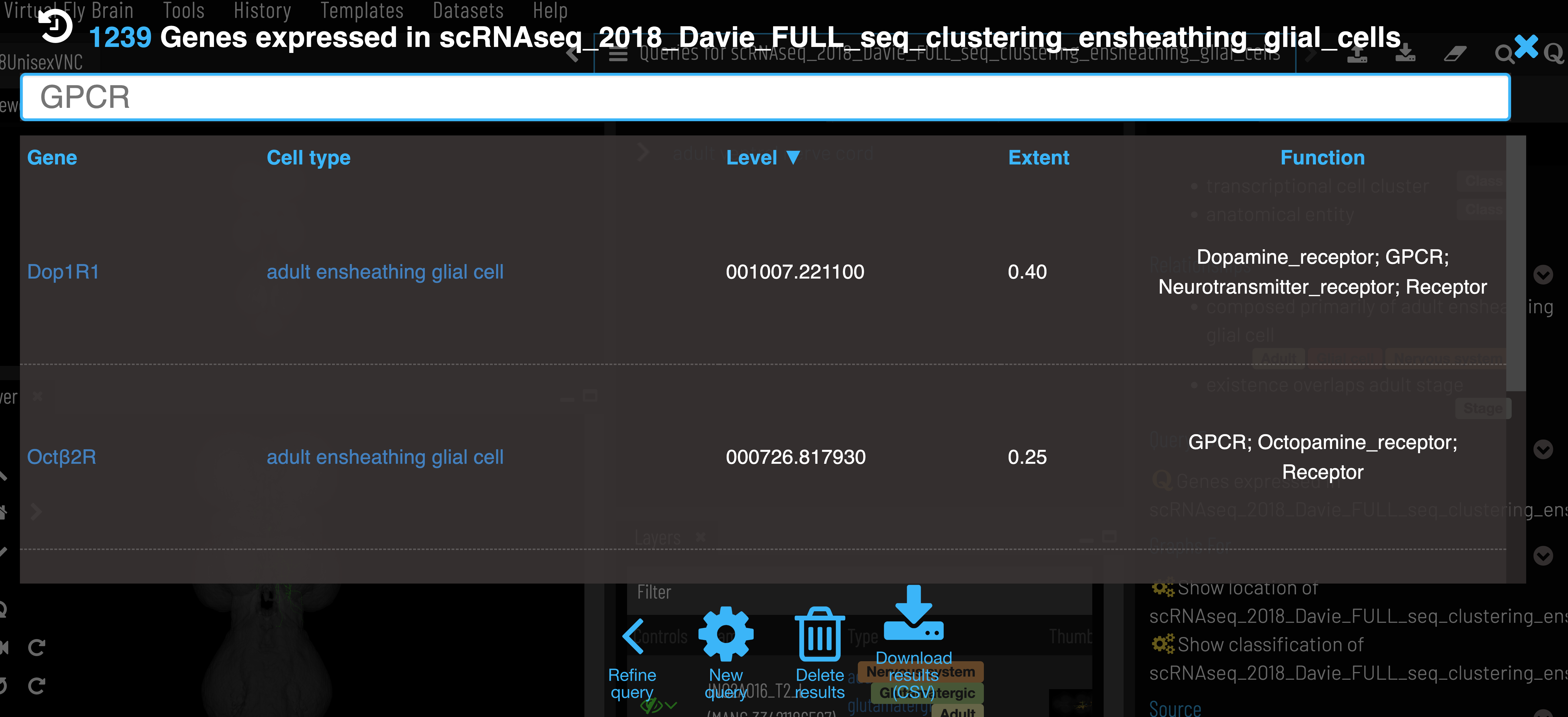
Expression level is the mean counts per million reads of all cells in the cell type cluster that express the given gene. Expression extent is the proportion of cells within the cluster that express the given gene. VFB only shows genes that are expressed in at least 20% of cells in the cluster (Extent > 0.2).
Gene semantic tags
We add semantic tags to genes to allow quick searching and filtering of results (‘Function’ column of gene expression data). These tags are based on FlyBase Gene Group membership and GO annotations, which are all sourced from FlyBase. A full list of gene tags with their associated Gene group and GO terms can be found here.
API
Data can also be retrieved using VFB_connect.
Data sources and processing
- Raw scRNAseq data is ingested by the EBI Single Cell Expression Atlas (SCEA). Where possible, cells are linked to cell types from the Drosophila Anatomy Ontology based on author annotations. Data is reprocessed, filtered and reclustered.
- FlyBase takes data from EBI, keeping only cells that are linked to ontology terms and generating summary expression data for each cell type (cluster).
- VFB takes control/wild type expression data from FlyBase for datasets where at least one nervous system cell type is present and filter out any genes expressed in less than 20% of cells per cluster.
Database schema
A simplified schema showing how scRNAseq data is stored in the VFB Neo4j database is shown below:
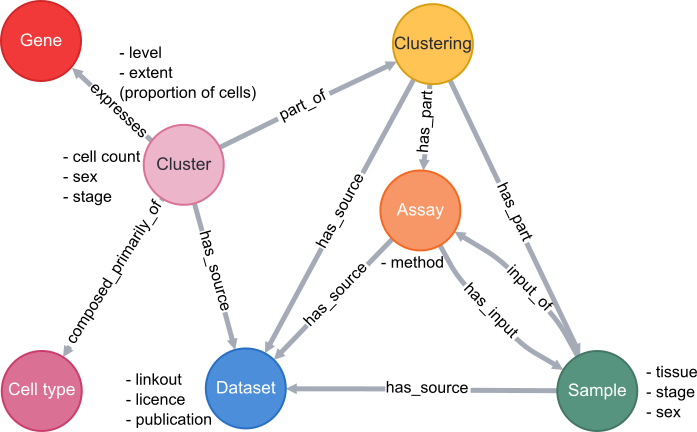
Cell types are classes from the Drosophila Anatomy Ontology and are also linked to several other types of data in VFB, such as images, connectomics and driver expression information.