This is the multi-page printable view of this section. Click here to print.
Tutorials
- 1: VFB Model Context Protocol (MCP) Tool Guide
- 2: Website Tutorials
- 3: Application Programming Interface (API) Tutorials
- 3.1: VFB connect API overview
- 3.2: Guide to Working with Images from Virtual Fly Brain (VFB) Using the VFBConnect Library
- 3.3: Downloading Images from VFB Using VFBconnect
- 3.4: Programmatic search using SOLR
- 3.5: Exploring Neurons in Navis
- 3.6: Plotting Neurons with Navis
- 3.7: pymaid
- 3.8: NBLAST
- 3.9: neuprint
1 - VFB Model Context Protocol (MCP) Tool Guide
Overview
The Virtual Fly Brain Model Context Protocol (MCP) Tool enables you to query VFB data through Large Language Models like Claude using natural language. This guide shows you how to get started and provides examples of common queries.
What is MCP?
The Model Context Protocol is a standard that allows LLMs to interact with external data sources and tools. The VFB MCP tool follows this standard, providing your LLM with access to VFB’s neuroanatomical databases, NBLAST similarity scores, and term information.
Accessing the Tool
The VFB MCP tool is available at: vfb3-mcp.virtualflybrain.org
Quick Start
Use the Live Service (Recommended)
The easiest way to use VFB3-MCP is through our hosted service. This requires no installation or setup on your machine.
Claude Desktop Setup
- Open Claude Desktop and go to Settings
- Navigate to the MCP section
- Add a new MCP server with these settings:
- Server Name:
virtual-fly-brain(or any name you prefer) - Type: HTTP
- Server URL:
https://vfb3-mcp.virtualflybrain.org
- Server Name:
Configuration JSON (alternative method):
{
"mcpServers": {
"virtual-fly-brain": {
"type": "http",
"url": "https://vfb3-mcp.virtualflybrain.org",
"tools": ["*"]
}
}
}
Claude Code Setup
- Locate your Claude configuration file:
- macOS/Linux:
~/.claude.json - Windows:
%USERPROFILE%\.claude.json
- macOS/Linux:
- Add the VFB3-MCP server to your configuration:
{
"mcpServers": {
"virtual-fly-brain": {
"type": "http",
"url": "https://vfb3-mcp.virtualflybrain.org",
"tools": ["*"]
}
}
}
- Restart Claude Code for changes to take effect
GitHub Copilot Setup
- Open VS Code with GitHub Copilot installed
- Open Settings (
Ctrl/Cmd + ,) - Search for “MCP” in the settings search
- Find the MCP Servers setting
- Add the server URL:
https://vfb3-mcp.virtualflybrain.org - Give it a name like “Virtual Fly Brain”
Alternative JSON configuration (in mcp.json):
{
"servers": {
"virtual-fly-brain": {
"type": "http",
"url": "https://vfb3-mcp.virtualflybrain.org"
}
}
}
Visual Studio Code (with MCP Extension)
- Install the MCP extension for VS Code from the marketplace
- Open the Command Palette (
Ctrl/Cmd + Shift + P) - Type “MCP: Add server” and select it
- Choose “HTTP” as the server type
- Enter the server details:
- Name:
virtual-fly-brain - URL:
https://vfb3-mcp.virtualflybrain.org
- Name:
- Save and restart VS Code if prompted
Other MCP Clients
For any MCP-compatible client that supports HTTP servers:
{
"mcpServers": {
"virtual-fly-brain": {
"type": "http",
"url": "https://vfb3-mcp.virtualflybrain.org",
"tools": ["*"]
}
}
}
Gemini Setup
To use the Virtual Fly Brain (VFB) Model Context Protocol (MCP) server with Google Gemini, you can connect through custom Python/Node.js clients that support MCP.
Note: Direct Gemini web interface integration with MCP is not currently supported. Developer tools are needed to connect the two.
Option 1: Using Python
For application development, use the mcp and google-genai libraries to connect.
Setup: pip install google-genai mcp
Implementation: Use an SSEClientTransport to connect to the VFB URL, list its tools, and pass their schemas to the Gemini model as Function Declarations.
Testing the Connection
Once configured, you can test that VFB3-MCP is working by asking your AI assistant questions like:
Basic Queries:
- “Get information about the neuron VFB_jrcv0i43”
- “Search for terms related to medulla in the fly brain”
- “What neurons are in the antennal lobe?”
Advanced Queries:
- “Find all neurons that connect to the mushroom body”
- “Show me expression patterns for gene repo”
- “What brain regions are involved in olfactory processing?”
- “Run a connectivity analysis for neuron VFB_00101567”
Search Examples:
- “Search for adult neurons in the visual system”
- “Find genes expressed in the central complex”
- “Show me all templates available in VFB”
If you see responses with VirtualFlyBrain data, including neuron names, brain regions, gene expressions, or connectivity information, the setup is successful!
For more detailed usage examples and API calls, see examples.md.
Local Installation
If you prefer to run the MCP server locally, see the VFB3-MCP repository README for detailed installation instructions.
Core Features
The tool provides access to three main capabilities:
1. Term Information Queries (get_term_info)
Retrieve detailed information about any VFB term using its VFB ID.
Example Query:
"What is the medulla? Please get the full definition and structure."
Returns:
- Term definition and synonyms
- Classification and type information
- Anatomical relationships (part of, develops from, innervates, etc.)
- Associated neurons and expression patterns
- Related images and connectivity data
2. Term Search (search_terms)
Search for VFB terms using keywords and filters.
Example Query:
"Find neurons in the medulla"
Advanced Filtering Options:
- Filter by entity type: neuron, muscle, glia, anatomical region
- Filter by nervous system component: visual system, olfactory system, sensory neuron, motor neuron, etc.
- Filter by nervous system property: cholinergic, GABAergic, glutamatergic, dopaminergic, peptidergic, etc.
- Filter by dataset: FAFB, FlyCircuit, hemibrain, neuprint, flycircuit, etc.
3. Query Execution (run_query)
Execute specific queries on VFB terms, including NBLAST similarity analysis.
Example Query:
"What neurons are morphologically similar to IN02A049?"
Example Use Cases
Case 1: Exploring a Transgenic Construct
User Question: “What is P{E(spl)m8-HLH-2.61} and where is it used?”
Tool Actions:
- Search for the construct in VFB
- Retrieve full term information
- Return detailed description including:
- FlyBase ID (FBtp0004163)
- Gene involved: E(spl)m8
- Type: Transgenic construct (reporter for gene expression)
- Expression patterns and research use
- Related constructs and variants
Result: You get comprehensive information about the transgenic reporter, its purpose, and research applications.
Case 2: Understanding a Neuroblast Population
User Question: “What is the Medulla Forming Neuroblast and what role does it play?”
Tool Actions:
- Search for “medulla forming neuroblast”
- Get term info for FBbt_00001938
- Return:
- Cell type classification (neuroblast)
- Location in the larval optic anlage
- Developmental fate (produces medulla neurons)
- Marker genes (dpn, ase)
- Number of neurons produced (~54,000 secondary neurons)
- Available scRNAseq and expression data
Result: Understand the neuroblast’s role in developing the adult medulla’s neural circuitry.
Case 3: Discovering Neuron Types
User Question: “What types of neurons are in the medulla?”
Tool Actions:
- Search for “neuron” with filter for medulla
- Return 472 neuron types with parts in the medulla, organized by category:
- Medulla intrinsic neurons (Mi): columnar neurons with confined processes
- Central medulla intrinsic neurons (Cm): arborizing in central/serpentine layer
- Medulla tangential neurons (Mt): wide-field spanning neurons
- Medulla visual projection neurons (MeVP): tangential projection neurons
- Medulla columnar neurons (MC): connecting medulla to tubercle
- Plus many more specialized types
Result: Get a comprehensive overview of medulla neuron diversity and organization.
Case 4: Morphological Similarity Queries
User Question: “Show me what the IN02A049 neuron looks like and find similar neurons.”
Tool Actions:
- Get term info for IN02A049 (specific instance from MANC connectome)
- Execute NBLAST query for morphologically similar neurons
- Return:
- 3D visualization of the neuron
- Classification and properties
- Synaptic inputs (neurotransmitter types and counts)
- Morphologically similar neurons with NBLAST scores
- Cross-connectome comparisons
Result: Discover morphologically similar neurons for comparative analysis.
Case 5: Understanding NBLAST Scores
User Question: “How are NBLAST scores calculated?”
Tool Actions:
- Search for “NBLAST”
- Return detailed explanation including:
- Algorithm basis (Costa et al. 2016)
- How neurons are represented (point skeletons with direction vectors)
- Scoring mechanism (Euclidean distance + direction similarity)
- Normalization approach (comparison to self-match)
- Asymmetry property and symmetrical variants
- Available datasets (FAFB-FlyWire, Male-CNS optic lobe, FlyCircuit, Hemibrain, FAFB-CATMAID)
Result: Understand the methodology behind VFB’s morphological similarity analysis.
Tips for Effective Queries
1. Use VFB IDs When Known
If you know the VFB ID of a term, use it directly:
"Get me detailed information about FBbt_00003748 (the medulla)"
2. Combine Search and Query
Use search first to find relevant terms, then query them:
"Find all cholinergic neurons in the visual system,
then tell me about the top 5"
3. Filter Strategically
Use filters to narrow results:
"Show me motor neurons from the male-CNS optic lobe dataset"
4. Explore Relationships
Ask about anatomical and functional relationships:
"What neurons innervate the medulla?
What brain regions do they come from?"
5. Cross-Dataset Comparisons
Compare neurons across different connectome datasets:
"Find the equivalent of this FAFB neuron in the hemibrain dataset"
Available Datasets
The VFB MCP tool provides access to neurons and data from:
- FAFB-FlyWire (v783) - Large-scale adult brain connectome
- Male-CNS optic lobe (v1.0.1) - Focused optic lobe connectome
- FlyCircuit (1.0) - Single-neuron morphology database
- Hemibrain (1.2.1) - Subset of adult brain connectome
- FAFB-CATMAID - Manually traced EM data
- FlyLight Split-GAL4 - Driver line expression images
- scRNAseq data - Transcriptomic information
Understanding VFB IDs
VFB terms use standardized identifiers:
- FBbt_ - Drosophila anatomy terms (e.g., FBbt_00003748 = medulla)
- FBgn_ - FlyBase genes (e.g., FBgn0000137 = ase gene)
- FBtp_ - FlyBase transgenic constructs (e.g., FBtp0004163 = P{E(spl)m8-HLH-2.61})
- VFB_ - VirtualFlyBrain-specific IDs for individuals and images
Next Steps
- Explore the tool: Visit vfb3-mcp.virtualflybrain.org
- Read more about NBLAST: See our NBLAST concepts page
- Learn about VFB APIs: Check our API overview
- Browse VFB directly: Visit the main Virtual Fly Brain site
Common Questions
Q: Do I need to install anything? A: No, if you’re using Claude or another MCP-compatible LLM with the integration already set up, just start asking questions.
Q: Can I download the data I find? A: Yes, many results include links to download images, neuron skeletons, and connectivity data. Check the API documentation for programmatic access.
Q: What if I don’t know the VFB ID? A: The search tool can find terms by name or keyword. The LLM will help you locate the right term.
Q: Can I combine VFB queries with other analyses? A: Absolutely! You can ask your LLM to retrieve VFB data and then perform additional analyses, create visualizations, or integrate with other tools.
Happy exploring! If you have questions or suggestions about the VFB MCP tool, please reach out to the VFB team.
2 - Website Tutorials
2.1 - Similarity Score Queries Guide
Scores Availability
The similarity scores are calculated using NBLAST or other third-party scores (e.g. NeuronBridge). Only queries with above-threshold scores will appear in the ‘Find similar…’ menu.
Step 1: Select a Neuron or Expression Pattern
Navigate to the VFB browser and select a neuron or expression pattern of interest so it’s information is shown in the Term Info panel.
Step 2: Open the ‘Find similar…’ Menu under the ‘Query For’ section
Under the ‘Query For’ section, locate the ‘Find similar…’ expandable menu. This menu will only appear if an above-threshold score is available for the selected neuron or expression pattern.
Step 3: Run a Similarity Score Query
Inside the ‘Find similar…’ menu, you will find queries to find morphologically similar neurons or expression patterns. Select a query to run it. The results will display neurons or expression patterns that are similar to your selected neuron or expression pattern.

Note: We recalculate these scores shortly after new data is added, so new matches may appear if you check back a few months later.
2.2 - Bulk Image Download
Using the Bulk Image Download Tool
Follow these steps to download multiple files from VirtualFlyBrain using the bulk download tool:
Step 1: Open the Images
Navigate to the VFB Browser and open all the images you are interested in downloading. You can do this by searching for specific images and opening their respective pages.
Step 2: Open the Bulk Download Tool
Once you have all the images open, locate the bulk download icon in the top right. Click on this icon to open the bulk download tool.
Step 3: Select the Data Type
In the bulk download tool, select the type of data you want to download. The options are OBJ, SWC, NRRD, and References.
Step 4: Select the Variables
If applicable, select specific variables related to the data type you have chosen.
Step 5: Download the Data
After making your selections, click the ‘Download’ button to start the download. The data will be downloaded in a zip file named “VFB Files.zip”.
Step 6: Unzip the File
Once the download is complete, locate the zip file in your downloads folder and unzip it to access the data.
Step 7: Open the Data
Finally, open the data in your chosen software. For example, if you chose OBJ, you could open the data in a 3D modeling program.
Note: If something goes wrong during the download, an error message will be displayed and you will be prompted to try again. If no entries are found for the selected types and variables, a message will be displayed to inform you.
2.3 - Browsing single cell RNAseq data
Finding transcriptomics data on VFB
Finding cell type clusters
Cell types with available scRNAseq data can be identified using the ‘Has scRNAseq data filter’ when searching.
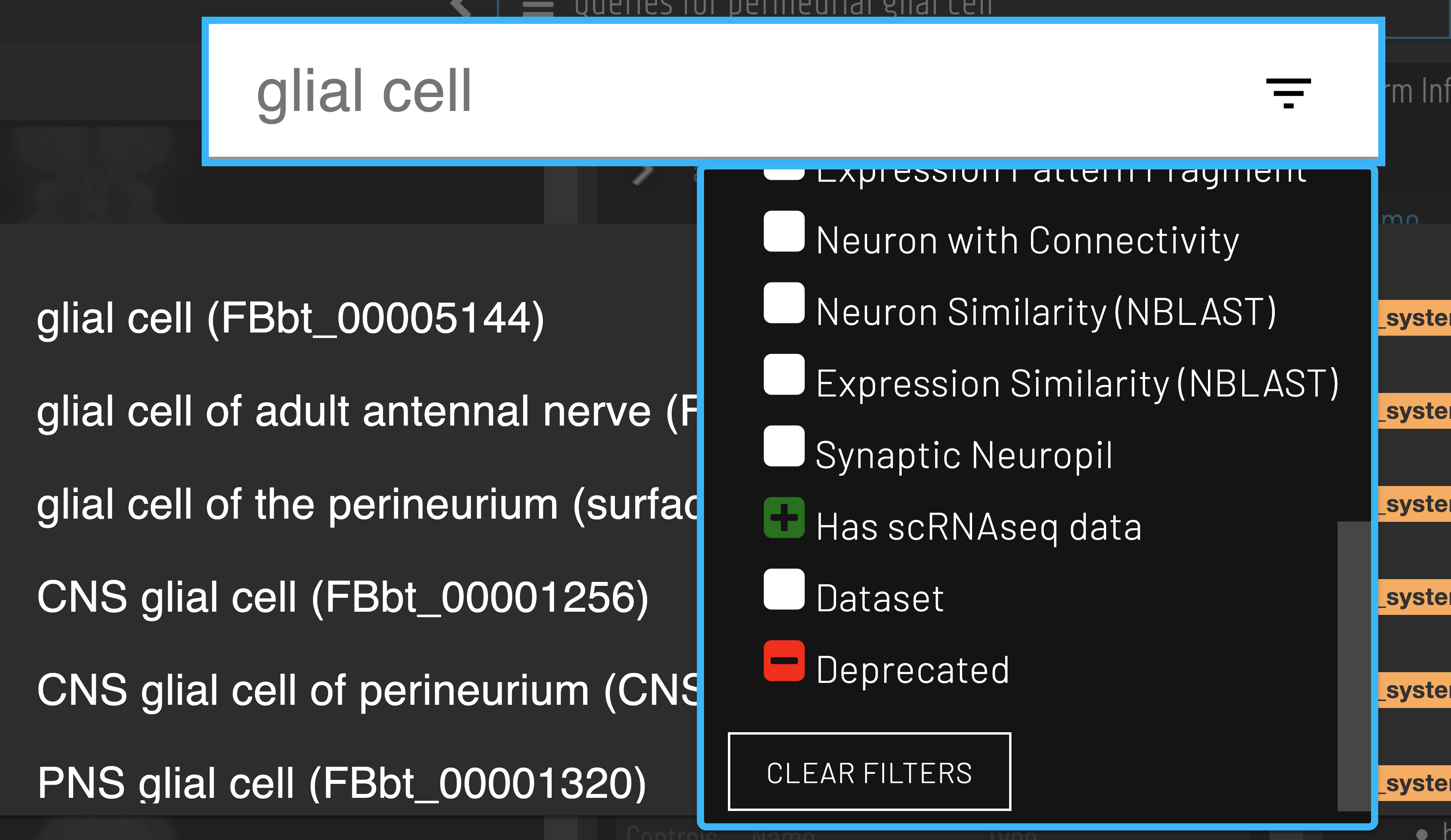
Clusters for a cell type of interest can be found via the ‘Single cell transcriptomics data for…’ query.

Gene expression data
After selecting a cluster, gene expression can be retrieved via the ‘Genes expressed in…’ query.
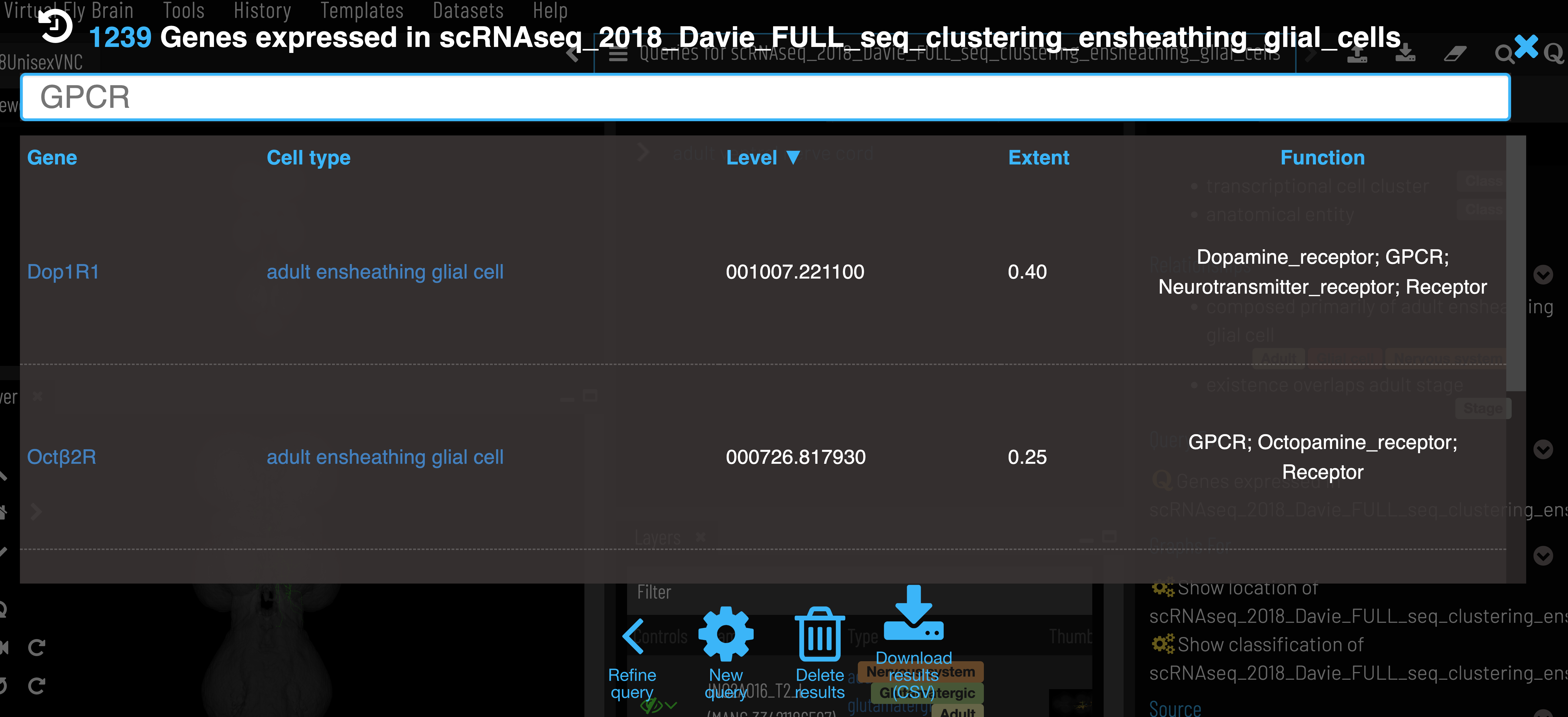
Expression level is the mean counts per million reads of all cells in the cell type cluster that express the given gene. Expression extent is the proportion of cells within the cluster that express the given gene. VFB only shows genes that are expressed in at least 20% of cells in the cluster (Extent > 0.2).
Gene semantic tags
We add semantic tags to genes to allow quick searching and filtering of results (‘Function’ column of gene expression data). These tags are based on FlyBase Gene Group membership and GO annotations, which are all sourced from FlyBase. A full list of gene tags with their associated Gene group and GO terms can be found here.
API
Data can also be retrieved using VFB_connect.
Data sources and processing
- Raw scRNAseq data is ingested by the EBI Single Cell Expression Atlas (SCEA). Where possible, cells are linked to cell types from the Drosophila Anatomy Ontology based on author annotations. Data is reprocessed, filtered and reclustered.
- FlyBase takes data from EBI, keeping only cells that are linked to ontology terms and generating summary expression data for each cell type (cluster).
- VFB takes control/wild type expression data from FlyBase for datasets where at least one nervous system cell type is present and filter out any genes expressed in less than 20% of cells per cluster.
Database schema
A simplified schema showing how scRNAseq data is stored in the VFB Neo4j database is shown below:
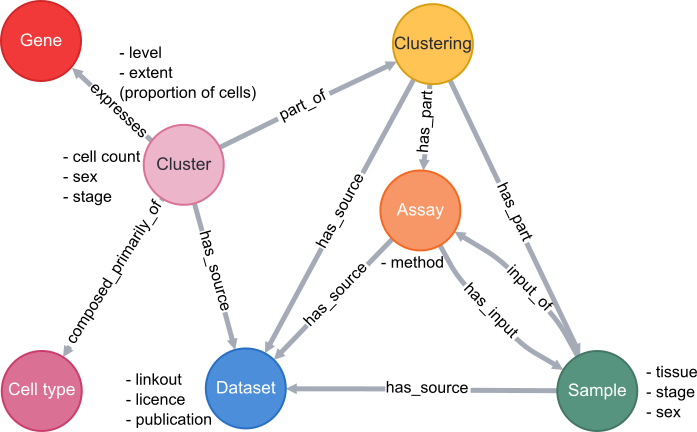
Cell types are classes from the Drosophila Anatomy Ontology and are also linked to several other types of data in VFB, such as images, connectomics and driver expression information.
3 - Application Programming Interface (API) Tutorials
3.1 - VFB connect API overview
VFB connect API overview
The VFB connect API provides programmatic access to the databases underlying Virtual Fly Brain.
At the core of Virtual Fly Brain is a set of curated terms for Drosophila neuro-anatomy organised into a queryable classification, including terms for brain regions, e.g. nodulus and neurons e.g. MBON01. These terms are used to annotate and classify individual brain regions and neurons in images and connectomics data. For example the term MBON01 is used to classify individual neurons from sources including the CATMAID-FAFB and Neuprint-HemiBrain databases. VFB stores both registered 3D images and connectomics data (where available) for all of these neurons.
A single VfbConnect object wraps database connections and canned queries against all open VFB databases. It includes methods for retreiving metadata about anatomy, individual brain regions and neurons including IDs for these that can be used for queries against other databases (e.g. CATMAID & neuprint). It provides methods for downloading images and connectomics data. It provides access to sophisticated queries for anatomical classes and individual neurons according to their classification & properties.
Locations for methods under a VfbConnect object.
- Under
vc.neo_query_wrapperare- A set of methods that take lists of IDs as a primary argument and return metadata.
- A set of methods for mapping between VFB IDs and external IDs
- Directly under
vcare:- A set of methods that take the names of classes in VFB e.g. ’nodulus’ or ‘Kenyon cell’, or simple query expressions using the names of classes and return metadata about the classes.
- A set methods for querying connectivity and similarity
- Direct access to API queries is provided under the ’nc’ and ‘oc’ attributes for Neo4J and OWL queries respectively. We will not cover details of how to use these here.
Note: available methods and their documentation are easy to explore in DeepNote. Tab completion and type adhead can be used to help find methods. Float your cursor over a method to see its signature and docstring.
1. vc.neo_query_wrapper methods overview
1.1 vc.neo_query_wrapper TermInfo queries return the results of a VFB Term Information window as JSON, following the VFB_JSON standard, or a summary that can easily be converted into a DataFrame.
# A query for full TermInfo. This probably produces more information than you will need for most purposes.
vc.neo_query_wrapper.get_type_TermInfo(['FBbt_00003686'])
[{'term': {'core': {'iri': 'http://purl.obolibrary.org/obo/FBbt_00003686',
'symbol': '',
'types': ['Entity',
'Anatomy',
'Nervous_system',
'Cell',
'Neuron',
'Class'],
'label': 'Kenyon cell',
'short_form': 'FBbt_00003686'},
'description': ['Intrinsic neuron of the mushroom body. They have tightly-packed cell bodies, situated in the rind above the calyx of the mushroom body (Ito et al., 1997). Four short fascicles, one per lineage, extend from the cell bodies of the Kenyon cells into the calyx (Ito et al., 1997). These 4 smaller fascicles converge in the calyx where they arborize and form pre- and post-synaptic terminals (Christiansen et al., 2011), with different Kenyon cells receiving input in different calyx regions/accessory calyces (Tanaka et al., 2008). They emerge from the calyx as a thick axon bundle referred to as the peduncle that bifurcates to innervate the dorsal and medial lobes of the mushroom body (Tanaka et al., 2008).'],
'comment': ['Pre-synaptic terminals were identified using two presynaptic markers (Brp and Dsyd-1) and post-synaptic terminals by labelling a subunit of the acetylcholine receptor (Dalpha7) in genetically labelled Kenyon cells (Christiansen et al., 2011).']},
'query': 'Get JSON for Class',
'version': '44725ae',
'parents': [{'iri': 'http://purl.obolibrary.org/obo/FBbt_00001366',
'symbol': '',
'types': ['Entity',
'Anatomy',
'Nervous_system',
'Cell',
'Neuron',
'Class'],
'label': 'supraesophageal ganglion neuron',
'short_form': 'FBbt_00001366'},
{'iri': 'http://purl.obolibrary.org/obo/FBbt_00007484',
'symbol': '',
'types': ['Entity',
'Anatomy',
'Nervous_system',
'Cell',
'Neuron',
'Class'],
'label': 'mushroom body intrinsic neuron',
'short_form': 'FBbt_00007484'}],
'relationships': [{'relation': {'type': 'develops_from',
'iri': 'http://purl.obolibrary.org/obo/RO_0002202',
'label': 'develops from'},
'object': {'iri': 'http://purl.obolibrary.org/obo/FBbt_00007113',
'symbol': '',
'types': ['Entity',
'Anatomy',
'Nervous_system',
'Cell',
'Neuroblast',
'Class'],
'label': 'mushroom body neuroblast',
'short_form': 'FBbt_00007113'}},
{'relation': {'type': 'overlaps',
'iri': 'http://purl.obolibrary.org/obo/RO_0002131',
'label': 'overlaps'},
'object': {'iri': 'http://purl.obolibrary.org/obo/FBbt_00003687',
'symbol': '',
'types': ['Entity',
'Synaptic_neuropil',
'Anatomy',
'Nervous_system',
'Synaptic_neuropil_domain',
'Class'],
'label': 'mushroom body pedunculus',
'short_form': 'FBbt_00003687'}},
{'relation': {'type': 'part_of',
'iri': 'http://purl.obolibrary.org/obo/BFO_0000050',
'label': 'is part of'},
'object': {'iri': 'http://purl.obolibrary.org/obo/FBbt_00005801',
'symbol': '',
'types': ['Entity',
'Synaptic_neuropil',
'Anatomy',
'Nervous_system',
'Synaptic_neuropil_block',
'Class'],
'label': 'mushroom body',
'short_form': 'FBbt_00005801'}},
{'relation': {'type': 'receives_synaptic_input_in',
'iri': 'http://purl.obolibrary.org/obo/RO_0013002',
'label': 'receives synaptic input in'},
'object': {'iri': 'http://purl.obolibrary.org/obo/FBbt_00003685',
'symbol': '',
'types': ['Entity',
'Synaptic_neuropil',
'Anatomy',
'Nervous_system',
'Synaptic_neuropil_domain',
'Class'],
'label': 'mushroom body calyx',
'short_form': 'FBbt_00003685'}}],
'xrefs': [],
'anatomy_channel_image': [{'channel_image': {'channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_jrchjwig',
'symbol': '',
'types': ['Entity', 'Individual'],
'label': 'KCg-t_R - 5812981989_c',
'short_form': 'VFBc_jrchjwig'},
'image': {'template_channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_00101567',
'symbol': '',
'types': ['Entity', 'Individual', 'Template'],
'label': 'JRC2018Unisex_c',
'short_form': 'VFBc_00101567'},
'index': [],
'template_anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_00101567',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'Nervous_system',
'Individual',
'Template'],
'label': 'JRC2018Unisex',
'short_form': 'VFB_00101567'},
'image_folder': 'http://www.virtualflybrain.org/data/VFB/i/jrch/jwig/VFB_00101567/'},
'imaging_technique': {'iri': 'http://purl.obolibrary.org/obo/FBbi_00050000',
'symbol': 'FIB-SEM',
'types': ['Entity', 'Class'],
'label': 'focussed ion beam scanning electron microscopy (FIB-SEM)',
'short_form': 'FBbi_00050000'}},
'anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_jrchjwig',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'has_neuron_connectivity',
'Cell',
'Individual',
'has_region_connectivity',
'NBLAST',
'Nervous_system',
'Neuron'],
'label': 'KCg-t_R - 5812981989',
'short_form': 'VFB_jrchjwig'}},
{'channel_image': {'channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_jrchjwig',
'symbol': '',
'types': ['Entity', 'Individual'],
'label': 'KCg-t_R - 5812981989_c',
'short_form': 'VFBc_jrchjwig'},
'image': {'template_channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_00101384',
'symbol': '',
'types': ['Entity', 'Individual', 'Template'],
'label': 'JRC_FlyEM_Hemibrain_c',
'short_form': 'VFBc_00101384'},
'index': [],
'template_anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_00101384',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'Nervous_system',
'Individual',
'Template'],
'label': 'JRC_FlyEM_Hemibrain',
'short_form': 'VFB_00101384'},
'image_folder': 'http://www.virtualflybrain.org/data/VFB/i/jrch/jwig/VFB_00101384/'},
'imaging_technique': {'iri': 'http://purl.obolibrary.org/obo/FBbi_00050000',
'symbol': 'FIB-SEM',
'types': ['Entity', 'Class'],
'label': 'focussed ion beam scanning electron microscopy (FIB-SEM)',
'short_form': 'FBbi_00050000'}},
'anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_jrchjwig',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'has_neuron_connectivity',
'Cell',
'Individual',
'has_region_connectivity',
'NBLAST',
'Nervous_system',
'Neuron'],
'label': 'KCg-t_R - 5812981989',
'short_form': 'VFB_jrchjwig'}},
{'channel_image': {'channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_jrchjwih',
'symbol': '',
'types': ['Entity', 'Individual'],
'label': 'KCg-t_R - 1392655948_c',
'short_form': 'VFBc_jrchjwih'},
'image': {'template_channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_00101567',
'symbol': '',
'types': ['Entity', 'Individual', 'Template'],
'label': 'JRC2018Unisex_c',
'short_form': 'VFBc_00101567'},
'index': [],
'template_anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_00101567',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'Nervous_system',
'Individual',
'Template'],
'label': 'JRC2018Unisex',
'short_form': 'VFB_00101567'},
'image_folder': 'http://www.virtualflybrain.org/data/VFB/i/jrch/jwih/VFB_00101567/'},
'imaging_technique': {'iri': 'http://purl.obolibrary.org/obo/FBbi_00050000',
'symbol': 'FIB-SEM',
'types': ['Entity', 'Class'],
'label': 'focussed ion beam scanning electron microscopy (FIB-SEM)',
'short_form': 'FBbi_00050000'}},
'anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_jrchjwih',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'has_neuron_connectivity',
'Cell',
'Individual',
'has_region_connectivity',
'NBLAST',
'Nervous_system',
'Neuron'],
'label': 'KCg-t_R - 1392655948',
'short_form': 'VFB_jrchjwih'}},
{'channel_image': {'channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_jrchjwih',
'symbol': '',
'types': ['Entity', 'Individual'],
'label': 'KCg-t_R - 1392655948_c',
'short_form': 'VFBc_jrchjwih'},
'image': {'template_channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_00101384',
'symbol': '',
'types': ['Entity', 'Individual', 'Template'],
'label': 'JRC_FlyEM_Hemibrain_c',
'short_form': 'VFBc_00101384'},
'index': [],
'template_anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_00101384',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'Nervous_system',
'Individual',
'Template'],
'label': 'JRC_FlyEM_Hemibrain',
'short_form': 'VFB_00101384'},
'image_folder': 'http://www.virtualflybrain.org/data/VFB/i/jrch/jwih/VFB_00101384/'},
'imaging_technique': {'iri': 'http://purl.obolibrary.org/obo/FBbi_00050000',
'symbol': 'FIB-SEM',
'types': ['Entity', 'Class'],
'label': 'focussed ion beam scanning electron microscopy (FIB-SEM)',
'short_form': 'FBbi_00050000'}},
'anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_jrchjwih',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'has_neuron_connectivity',
'Cell',
'Individual',
'has_region_connectivity',
'NBLAST',
'Nervous_system',
'Neuron'],
'label': 'KCg-t_R - 1392655948',
'short_form': 'VFB_jrchjwih'}},
{'channel_image': {'channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_jrchjwii',
'symbol': '',
'types': ['Entity', 'Individual'],
'label': 'KCg-t_R - 785918963_c',
'short_form': 'VFBc_jrchjwii'},
'image': {'template_channel': {'iri': 'http://virtualflybrain.org/reports/VFBc_00101567',
'symbol': '',
'types': ['Entity', 'Individual', 'Template'],
'label': 'JRC2018Unisex_c',
'short_form': 'VFBc_00101567'},
'index': [],
'template_anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_00101567',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'Nervous_system',
'Individual',
'Template'],
'label': 'JRC2018Unisex',
'short_form': 'VFB_00101567'},
'image_folder': 'http://www.virtualflybrain.org/data/VFB/i/jrch/jwii/VFB_00101567/'},
'imaging_technique': {'iri': 'http://purl.obolibrary.org/obo/FBbi_00050000',
'symbol': 'FIB-SEM',
'types': ['Entity', 'Class'],
'label': 'focussed ion beam scanning electron microscopy (FIB-SEM)',
'short_form': 'FBbi_00050000'}},
'anatomy': {'iri': 'http://virtualflybrain.org/reports/VFB_jrchjwii',
'symbol': '',
'types': ['Entity',
'has_image',
'Adult',
'Anatomy',
'has_neuron_connectivity',
'Cell',
'Individual',
'has_region_connectivity',
'NBLAST',
'Nervous_system',
'Neuron'],
'label': 'KCg-t_R - 785918963',
'short_form': 'VFB_jrchjwii'}}],
'pub_syn': [{'pub': {'core': {'iri': 'http://flybase.org/reports/Unattributed',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': '',
'short_form': 'Unattributed'},
'FlyBase': '',
'PubMed': '',
'DOI': ''},
'synonym': {'type': '', 'label': 'KC', 'scope': 'has_exact_synonym'}},
{'pub': {'core': {'iri': 'http://flybase.org/reports/FBrf0236359',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': 'Eichler et al., 2017, Nature 548(7666): 175--182',
'short_form': 'FBrf0236359'},
'FlyBase': 'FBrf0236359',
'PubMed': '28796202',
'DOI': '10.1038/nature23455'},
'synonym': {'type': '',
'label': 'mature Kenyon cell',
'scope': 'has_exact_synonym'}},
{'pub': {'core': {'iri': 'http://flybase.org/reports/FBrf0111409',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': 'Lee et al., 1999, Development 126(18): 4065--4076',
'short_form': 'FBrf0111409'},
'FlyBase': '',
'PubMed': '10457015',
'DOI': ''},
'synonym': {'type': '',
'label': 'MB neuron',
'scope': 'has_narrow_synonym'}}],
'def_pubs': [{'core': {'iri': 'http://flybase.org/reports/FBrf0214059',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': 'Christiansen et al., 2011, J. Neurosci. 31(26): 9696--9707',
'short_form': 'FBrf0214059'},
'FlyBase': '',
'PubMed': '21715635',
'DOI': '10.1523/JNEUROSCI.6542-10.2011'},
{'core': {'iri': 'http://flybase.org/reports/FBrf0092568',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': 'Ito et al., 1997, Development 124(4): 761--771',
'short_form': 'FBrf0092568'},
'FlyBase': '',
'PubMed': '9043058',
'DOI': ''},
{'core': {'iri': 'http://flybase.org/reports/FBrf0205263',
'symbol': '',
'types': ['Entity', 'Individual', 'pub'],
'label': 'Tanaka et al., 2008, J. Comp. Neurol. 508(5): 711--755',
'short_form': 'FBrf0205263'},
'FlyBase': '',
'PubMed': '18395827',
'DOI': '10.1002/cne.21692'}]}]
# A query for summary info
import pandas as pd
summary = vc.neo_query_wrapper.get_type_TermInfo(['FBbt_00003686'], summary=True)
summary_tab = pd.DataFrame.from_records(summary)
summary_tab
| label | symbol | id | tags | parents_label | parents_id | |
|---|---|---|---|---|---|---|
| 0 | Kenyon cell | FBbt_00003686 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | supraesophageal ganglion neuron|mushroom body ... | FBbt_00001366|FBbt_00007484 |
# A different method is needed to get info about individual neurons
summary = vc.neo_query_wrapper.get_anatomical_individual_TermInfo(['VFB_jrchjrch'], summary=True)
summary_tab = pd.DataFrame.from_records(summary)
summary_tab
| label | symbol | id | tags | parents_label | parents_id | data_source | accession | templates | dataset | license | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 5-HTPLP01_R - 1324365879 | VFB_jrchjrch | Entity|has_image|Adult|Anatomy|has_neuron_conn... | adult serotonergic PLP neuron | FBbt_00110945 | neuprint_JRC_Hemibrain_1point1 | 1324365879 | JRC_FlyEM_Hemibrain|JRC2018Unisex | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... |
1.2 The neo_query_wrapper also includes methods for mapping between IDs from different sources.
# Some bodyIDs of HemiBrain neurons from the neuprint DataBase:
bodyIDs = [1068958652, 571424748, 1141631198]
vc.neo_query_wrapper.xref_2_vfb_id(map(str, bodyIDs)) # Note IDs must be strings
{'1068958652': [{'db': 'neuronbridge', 'vfb_id': 'VFB_jrchjwda'},
{'db': 'neuronbridge', 'vfb_id': 'VFB_jrch06r9'},
{'db': 'neuprint_JRC_Hemibrain_1point0point1', 'vfb_id': 'VFB_jrch06r9'},
{'db': 'neuprint_JRC_Hemibrain_1point1', 'vfb_id': 'VFB_jrchjwda'}],
'571424748': [{'db': 'neuronbridge', 'vfb_id': 'VFB_jrch06r6'},
{'db': 'neuronbridge', 'vfb_id': 'VFB_jrchjwct'},
{'db': 'neuprint_JRC_Hemibrain_1point0point1', 'vfb_id': 'VFB_jrch06r6'},
{'db': 'neuprint_JRC_Hemibrain_1point1', 'vfb_id': 'VFB_jrchjwct'}],
'1141631198': [{'db': 'neuronbridge', 'vfb_id': 'VFB_jrch05uz'},
{'db': 'neuronbridge', 'vfb_id': 'VFB_jrchjw8r'},
{'db': 'neuprint_JRC_Hemibrain_1point0point1', 'vfb_id': 'VFB_jrch05uz'},
{'db': 'neuprint_JRC_Hemibrain_1point1', 'vfb_id': 'VFB_jrchjw8r'}]}
# xref queries can be constrained by DB. Results can optionally be reversed
vc.neo_query_wrapper.xref_2_vfb_id(map(str, bodyIDs), db = 'neuprint_JRC_Hemibrain_1point1' , reverse_return=True)
{'VFB_jrchjw8r': [{'acc': '1141631198',
'db': 'neuprint_JRC_Hemibrain_1point1'}],
'VFB_jrchjwct': [{'acc': '571424748',
'db': 'neuprint_JRC_Hemibrain_1point1'}],
'VFB_jrchjwda': [{'acc': '1068958652',
'db': 'neuprint_JRC_Hemibrain_1point1'}]}
2. vc direct methods overview
2.1 Methods that take the names of classes in VFB e.g. ’nodulus’ or ‘Kenyon cell’, or simple query expressions using the names of classes and return metadata about the classes or individual neurons.
KC_types = vc.get_subclasses("Kenyon cell", summary=True)
pd.DataFrame.from_records(KC_types)
Running query: FBbt:00003686
Query URL: http://owl.virtualflybrain.org/kbs/vfb/subclasses?object=FBbt%3A00003686&prefixes=%7B%22FBbt%22%3A+%22http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FFBbt_%22%2C+%22RO%22%3A+%22http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FRO_%22%2C+%22BFO%22%3A+%22http%3A%2F%2Fpurl.obolibrary.org%2Fobo%2FBFO_%22%7D&direct=False
Query results: 37
| label | symbol | id | tags | parents_label | parents_id | |
|---|---|---|---|---|---|---|
| 0 | adult alpha'/beta' Kenyon cell | FBbt_00049834 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | alpha'/beta' Kenyon cell|adult Kenyon cell | FBbt_00100249|FBbt_00049825 | |
| 1 | immature Kenyon cell | FBbt_00047995 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | Kenyon cell | FBbt_00003686 | |
| 2 | gamma Kenyon cell | FBbt_00100247 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | Kenyon cell | FBbt_00003686 | |
| 3 | adult Kenyon cell | FBbt_00049825 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult MBp lineage neuron|Kenyon cell | FBbt_00110577|FBbt_00003686 | |
| 4 | gamma main Kenyon cell | KCg-m | FBbt_00111061 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | Kenyon cell of main calyx|adult gamma Kenyon cell | FBbt_00047926|FBbt_00049828 |
| 5 | alpha'/beta' anterior-posterior type 1 Kenyon ... | FBbt_00049859 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | alpha'/beta' anterior-posterior type 1 Kenyon ... | FBbt_00049836 | |
| 6 | alpha/beta Kenyon cell | FBbt_00100248 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | cholinergic neuron|adult Kenyon cell | FBbt_00007173|FBbt_00049825 | |
| 7 | gamma-s4 Kenyon cell | KCg-s4 | FBbt_00049832 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | gamma-s Kenyon cell | FBbt_00049830 |
| 8 | two-claw Kenyon cell | FBbt_00047997 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | multi-claw Kenyon cell | FBbt_00047994 | |
| 9 | single-claw Kenyon cell | FBbt_00047993 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | Kenyon cell | FBbt_00003686 | |
| 10 | alpha/beta posterior Kenyon cell | KCab-p | FBbt_00110931 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | alpha/beta Kenyon cell | FBbt_00100248 |
| 11 | alpha'/beta' middle Kenyon cell | KCa'b'-m | FBbt_00100253 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | Kenyon cell of main calyx|adult alpha'/beta' K... | FBbt_00047926|FBbt_00049834 |
| 12 | alpha/beta surface Kenyon cell | KCab-s | FBbt_00110930 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | alpha/beta surface/core Kenyon cell | FBbt_00049838 |
| 13 | alpha'/beta' anterior-posterior type 1 Kenyon ... | KCa'b'-ap1 | FBbt_00049836 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | alpha'/beta' anterior-posterior Kenyon cell | FBbt_00100250 |
| 14 | larval alpha'/beta' Kenyon cell | FBbt_00049835 | Entity|Neuron|Anatomy|Nervous_system|Cell|Larv... | larval Kenyon cell|alpha'/beta' Kenyon cell | FBbt_00049826|FBbt_00100249 | |
| 15 | gamma-s1 Kenyon cell | KCg-s1 | FBbt_00049787 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | gamma-s Kenyon cell | FBbt_00049830 |
| 16 | gamma-t Kenyon cell | KCg-t | FBbt_00049833 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult gamma Kenyon cell | FBbt_00049828 |
| 17 | four-claw Kenyon cell | FBbt_00047999 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | multi-claw Kenyon cell | FBbt_00047994 | |
| 18 | alpha'/beta' Kenyon cell | FBbt_00100249 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | Kenyon cell | FBbt_00003686 | |
| 19 | alpha/beta surface/core Kenyon cell | FBbt_00049838 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | Kenyon cell of main calyx|alpha/beta Kenyon cell | FBbt_00047926|FBbt_00100248 | |
| 20 | larval Kenyon cell | FBbt_00049826 | Entity|Neuron|Anatomy|Nervous_system|Cell|Larv... | Kenyon cell|embryonic/larval neuron | FBbt_00003686|FBbt_00001446 | |
| 21 | alpha'/beta' anterior-posterior Kenyon cell | FBbt_00100250 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult alpha'/beta' Kenyon cell | FBbt_00049834 | |
| 22 | six-claw Kenyon cell | FBbt_00048001 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | multi-claw Kenyon cell | FBbt_00047994 | |
| 23 | Kenyon cell of main calyx | FBbt_00047926 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult Kenyon cell | FBbt_00049825 | |
| 24 | alpha'/beta' anterior-posterior type 2 Kenyon ... | KCa'b'-ap2 | FBbt_00049837 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | alpha'/beta' anterior-posterior Kenyon cell|Ke... | FBbt_00100250|FBbt_00047926 |
| 25 | gamma-s3 Kenyon cell | KCg-s3 | FBbt_00049831 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | gamma-s Kenyon cell | FBbt_00049830 |
| 26 | alpha/beta inner-core Kenyon cell | FBbt_00049111 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | alpha/beta core Kenyon cell | FBbt_00110929 | |
| 27 | gamma dorsal Kenyon cell | KCg-d | FBbt_00110932 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult gamma Kenyon cell | FBbt_00049828 |
| 28 | alpha/beta outer-core Kenyon cell | FBbt_00049112 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | alpha/beta core Kenyon cell | FBbt_00110929 | |
| 29 | five-claw Kenyon cell | FBbt_00048000 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | multi-claw Kenyon cell | FBbt_00047994 | |
| 30 | alpha/beta core Kenyon cell | KCab-c | FBbt_00110929 | Entity|Neuron|Adult|Anatomy|Nervous_system|Cel... | alpha/beta surface/core Kenyon cell | FBbt_00049838 |
| 31 | multi-claw Kenyon cell | FBbt_00047994 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | Kenyon cell | FBbt_00003686 | |
| 32 | adult gamma Kenyon cell | FBbt_00049828 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | gamma Kenyon cell|adult Kenyon cell | FBbt_00100247|FBbt_00049825 | |
| 33 | three-claw Kenyon cell | FBbt_00047998 | Entity|Anatomy|Nervous_system|Cell|Neuron|Class | multi-claw Kenyon cell | FBbt_00047994 | |
| 34 | larval gamma Kenyon cell | FBbt_00049827 | Entity|Neuron|Anatomy|Nervous_system|Cell|Larv... | larval Kenyon cell|gamma Kenyon cell | FBbt_00049826|FBbt_00100247 | |
| 35 | gamma-s2 Kenyon cell | KCg-s2 | FBbt_00049788 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | gamma-s Kenyon cell | FBbt_00049830 |
| 36 | gamma-s Kenyon cell | FBbt_00049830 | Entity|Adult|Anatomy|Nervous_system|Cell|Neuro... | adult gamma Kenyon cell | FBbt_00049828 |
2.2 Methods for querying connectivity
Please see Connectivity Notebook for examples.
3.2 - Guide to Working with Images from Virtual Fly Brain (VFB) Using the VFBConnect Library
Prerequisites
Before starting, ensure you have the VFBConnect library installed. The recommended Python version is 3.10.14, as this version is tested against the library.
pip install vfb-connect
Importing the VFBConnect Library
Start by importing the VFBConnect library. This library provides a simple interface to interact with neuron data from the Virtual Fly Brain.
from vfb_connect import vfb
Retrieving Neuron Data
To work with specific neurons, you can use the vfb.term() function. This function takes a unique identifier (e.g., ID, label, synonym) for the neuron.
Example: Retrieving a Single Neuron
neuron = vfb.term('5th s-LNv (FlyEM-HB:511051477)')
The neuron variable now holds data about the neuron identified by the given term.
Working with Neuron Data
The retrieved neuron object can provide different representations of neuron data, such as its skeleton, mesh, and volume. These representations can be visualized using various plotting methods.
# Access the skeleton representation
neuron_skeleton = neuron.skeleton
# Check the type of the skeleton representation
print(type(neuron_skeleton)) # Output: <class 'navis.neuron.Neuron'>
# Plot the skeleton in 2D
neuron_skeleton.plot2d()
# Access the mesh representation
neuron_mesh = neuron.mesh
# Access the volume representation
neuron_volume = neuron.volume
Retrieving Multiple Neurons
You can retrieve multiple neurons using the vfb.terms() function, which accepts a list of neuron identifiers.
Example: Retrieving Multiple Neurons
neurons = vfb.terms(['5th s-LNv', 'fru-M-300008', 'catmaid_fafb:8876600'])
This command retrieves multiple neurons, which can then be visualized or manipulated collectively.
Flexible Matching Capabilities
One of the key features of the VFBConnect library is its flexible matching capability. The vfb.terms() function can accept a variety of identifiers, such as:
- IDs: Unique identifiers assigned to each neuron.
- Xref (Cross-references): External references that relate to other datasets.
- Labels: Human-readable names for neurons.
- Symbols: Abbreviated names or symbols used to represent neurons.
- Synonyms: Alternative names by which a neuron might be known.
- Partial Matching: You can provide a partial name, and VFBConnect will attempt to find the best match.
- Case Insensitive Matching: Matching is case insensitive, so if an exact match isn’t found, ‘5th s-LNv’ and ‘5TH S-LNV’ are treated the same. This allows for more flexible querying without worrying about exact case matching.
Example: Using Flexible Matching
neurons = vfb.terms('5th s-LN')
If an exact match isn’t found, VFBConnect will provide potential matches. This feature ensures that even with partial or approximate information, you can still retrieve the relevant neuron data.
Output Example:
Notice: No exact match found, but potential matches starting with '5th s-LN':
'5th s-LNv (FlyEM-HB:511051477)': 'VFB_jrchk8e0',
'5th s-LNv': 'VFB_jrchk8e0'
This notice will help you identify the correct neuron based on the closest matches.
Visualizing Neurons
VFBConnect provides various methods to visualize neuron data, both individually and collectively.
3D Visualization
To plot neurons in 3D, use the plot3d() method. This is useful for visualizing the spatial structure of neurons.
neurons.plot3d()
2D Visualization
For 2D visualization, use the plot2d() method.
neurons.plot2d()
Viewing Merged Templates
VFBConnect also allows viewing merged templates of neurons, combining multiple neuron structures into a single view.
neurons.show()
Opening Neurons in VFB
To open the neurons directly in Virtual Fly Brain, use the open() method. This will launch a browser window displaying the neurons in the VFB interface.
neurons.open()
Summary
- Use
vfb.term()to retrieve single neuron data. - Use
vfb.terms()to retrieve multiple neurons with support for partial, case-insensitive, and flexible matching (IDs, labels, symbols, synonyms, etc.). - Access different data representations (skeleton, mesh, volume) via neuron objects.
- Visualize neuron data in 2D and 3D.
- Use the
show()method to view merged neuron templates. - Open neuron data directly in Virtual Fly Brain with the
open()method.
These examples provide a foundation for working with neuron data from Virtual Fly Brain using the VFBConnect library. By exploring different neuron representations and visualization methods, you can analyze and understand neuron structures more effectively.
3.3 - Downloading Images from VFB Using VFBconnect
Introduction
VFBconnect is a Python package that provides an interface to the Virtual Fly Brain (VFB) API. It allows users to query the VFB database and download data, including images.
Installation
Before you can use VFBconnect, you need to install it. You can do this using pip:
pip install vfb-connect
Downloading Images
To download images from VFB using VFBconnect, you need to first import the package and create a client:
from vfb_connect.cross_server_tools import VfbConnect
vc = VfbConnect()
Next, you can use the get_images method to download images. This method requires the dataset ID as an argument:
dataset_id = 'your_dataset_id'
images = vc.get_images(dataset_id)
This will return a list of images from the specified dataset. Each image is represented as a dictionary with information such as the image ID, title, and URL.
To download the images, you can loop through the list and use the urlretrieve function from the urllib.request module:
import urllib.request
for image in images:
url = image['image_url']
filename = image['image_id'] + '.jpg'
urllib.request.urlretrieve(url, filename)
This will download each image and save it as a JPEG file in the current directory. The filename is the image ID.
Conclusion
This guide showed you how to use VFBconnect to download images from the Virtual Fly Brain based on a dataset. With VFBconnect, you can easily access and download data from VFB for your research.
3.4 - Programmatic search using SOLR
SOLR python example
an example using pysolr:
install:
pip install vfb-connect pysolr
example looking for label/name match:
import pysolr
solr = pysolr.Solr('https://solr.virtualflybrain.org/solr/ontology/')
term = 'medulla'
results = solr.search('label:"' + term + '"')
print(results.docs[0])
{'iri': ['http://purl.obolibrary.org/obo/FBbt_00003748'],
'obo_id_autosuggest': ['FBbt_00003748', 'FBbt:00003748', 'FBbt 00003748'],
'label_autosuggest': ['medulla', 'medulla', 'medulla'],
'synonym_autosuggest': ['ME', 'Med', 'optic medulla', 'm'],
'label': 'medulla',
'synonym': ['ME', 'Med', 'optic medulla', 'm'],
'short_form': 'FBbt_00003748',
'autosuggest': ['medulla', 'ME', 'Med', 'optic medulla', 'm'],
'facets_annotation': ['Entity',
'Adult',
'Anatomy',
'Class',
'Nervous_system',
'Synaptic_neuropil',
'Synaptic_neuropil_domain'],
'unique_facets': ['Nervous_system', 'Adult', 'Synaptic_neuropil_domain'],
'id': 'http://purl.obolibrary.org/obo/FBbt_00003748',
'shortform_autosuggest': ['FBbt_00003748', 'FBbt:00003748', 'FBbt 00003748'],
'obo_id': ['FBbt:00003748'],
'_version_': 1734360220689235970}
Note: any of the above fields can be searched (autosuggest being a combination of both label and synonyms)
3.5 - Exploring Neurons in Navis
TreeNeurons using navis.Overview
navis is a Python package for analysing, manipulating and visualizing neurons. Official documentation here.
Basic datatypes: neurons and neuron lists
navis knows three types of neurons:
TreeNeurons= skeletons, e.g. from CATMAIDMeshNeurons= meshes, e.g. from the hemibrain segmentationDotprops= points + tangent vectors (typically only used for NBLAST)
Collections of neurons are typically held in a specialized container: a NeuronList.
Neurons
In this notebook we will focus on skeletons - a.k.a. TreeNeurons - since this is what you get out of CATMAID. Let’s kick things off by having a look at what neurons look like once it’s loaded:
import navis
# Load one of the example neurons shipped with navis
# (these are olfactory projection neurons from the hemibrain data set)
n = navis.example_neurons(1, kind='skeleton')
# Print some basic info
n
WARNING: Could not load OpenGL library.
| type | navis.TreeNeuron |
|---|---|
| name | 1734350788 |
| id | 1734350788 |
| n_nodes | 4465 |
| n_connectors | None |
| n_branches | 603 |
| n_leafs | 619 |
| cable_length | 266457.994591 |
| soma | [4176] |
| units | 8 nanometer |
Above summary lists a couple of (computed) properties of the neuron. Each of those can also be accessed directly like so:
n.id
1734350788
There are many more properties that you might find interesting! Typing n. and pressing TAB should give auto-complete suggestions of available properties and methods. If your notebook editor has problems with that, you can fall back to using dir().
Here is an (incomplete) list of some of the more relevant properties:
bbox: bounding box of the neuroncable_length: cable lengthid: every neuron has an IDnodes: the SWC node table underlying the neuron
And some class methods:
reroot: reroot neuronplot2d/plot3d: plot the neuron (see also plotting turorial)copy: make and return a copyprune_twigs: remove small terminal twigs
As an example: this is how you get the ID of this neuron’s root node.
# Current root node of this neuron
n.root
array([1], dtype=int32)
Some of the properties such as .root or .ends are computed on-the-fly from the underlying raw data. For TreeNeurons that’s the node table (and its graph representation). The node table is a pandas DataFrame that looks effectively like a SWC:
# `.head()` gives us the first couple rows
n.nodes.head()
| node_id | label | x | y | z | radius | parent_id | type | |
|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | 15784.0 | 37250.0 | 28102.0 | 10.000000 | -1 | root |
| 1 | 2 | 0 | 15764.0 | 37230.0 | 28102.0 | 18.284300 | 1 | slab |
| 2 | 3 | 0 | 15744.0 | 37190.0 | 28142.0 | 34.721401 | 2 | slab |
| 3 | 4 | 0 | 15744.0 | 37150.0 | 28182.0 | 34.721401 | 3 | slab |
| 4 | 5 | 0 | 15704.0 | 37130.0 | 28242.0 | 34.721401 | 4 | slab |
The methods (such as .reroot) are short-hands for main navis functions:
# Reroot neuron to another node
n2 = n.reroot(new_root=2)
# Print the new root -> expect "2"
n2.root
array([2])
# Instead of calling the shorthand method, we can also do this
n3 = navis.reroot_neuron(n, new_root=2)
n3.root
array([2])
NeuronLists
In practice you will likely work with multiple neurons at a time. For that, navis has a convenient container: NeuronLists
# Get more than one example neuron
nl = navis.example_neurons(5)
# `nl` is a NeuronList
type(nl)
navis.core.neuronlist.NeuronList
# You can also create neuron lists yourself
my_nl = navis.NeuronList(n)
In many ways NeuronLists work like Python-lists with a couple of extras:
# Calling just the neuronlist produces a summary
nl
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 1734350788 | 1734350788 | 4465 | None | 603 | 619 | 266457.994591 | [4176] | 8 nanometer |
| 1 | navis.TreeNeuron | 1734350908 | 1734350908 | 4845 | None | 733 | 760 | 304277.007958 | [6] | 8 nanometer |
| 2 | navis.TreeNeuron | 722817260 | 722817260 | 4336 | None | 635 | 658 | 274910.568784 | None | 8 nanometer |
| 3 | navis.TreeNeuron | 754534424 | 754534424 | 4702 | None | 697 | 727 | 286742.998887 | [4] | 8 nanometer |
| 4 | navis.TreeNeuron | 754538881 | 754538881 | 4890 | None | 626 | 642 | 291434.992623 | [703] | 8 nanometer |
# Get a single neuron from the neuronlist
nl[1]
| type | navis.TreeNeuron |
|---|---|
| name | 1734350908 |
| id | 1734350908 |
| n_nodes | 4845 |
| n_connectors | None |
| n_branches | 733 |
| n_leafs | 760 |
| cable_length | 304277.007958 |
| soma | [6] |
| units | 8 nanometer |
neuronlists also support fancy indexing similar to numpy arrays:
# Get multiple neurons from the neuronlist
nl[[1, 2]]
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 1734350908 | 1734350908 | 4845 | None | 733 | 760 | 304277.007958 | [6] | 8 nanometer |
| 1 | navis.TreeNeuron | 722817260 | 722817260 | 4336 | None | 635 | 658 | 274910.568784 | None | 8 nanometer |
# Slicing is also supported
nl[1:3]
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 1734350908 | 1734350908 | 4845 | None | 733 | 760 | 304277.007958 | [6] | 8 nanometer |
| 1 | navis.TreeNeuron | 722817260 | 722817260 | 4336 | None | 635 | 658 | 274910.568784 | None | 8 nanometer |
Strings will be matched against the neurons’ names.
# Get neuron(s) by their name
nl['754534424']
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 754534424 | 754534424 | 4702 | None | 697 | 727 | 286742.998887 | [4] | 8 nanometer |
neuronlists have a special .idx indexer that let’s you select neurons by their ID
# Get neuron(s) by their ID
# -> note that for example neurons name == id
nl.idx[[754534424, 722817260]]
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 754534424 | 754534424 | 4702 | None | 697 | 727 | 286742.998887 | [4] | 8 nanometer |
| 1 | navis.TreeNeuron | 722817260 | 722817260 | 4336 | None | 635 | 658 | 274910.568784 | None | 8 nanometer |
# Access properties across neurons -> returns numpy arrays
nl.n_nodes
array([4465, 4845, 4336, 4702, 4890])
# Select neurons by given property
# -> this works with any boolean array
nl[nl.n_nodes >= 4500]
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | 1734350908 | 1734350908 | 4845 | None | 733 | 760 | 304277.007958 | [6] | 8 nanometer |
| 1 | navis.TreeNeuron | 754534424 | 754534424 | 4702 | None | 697 | 727 | 286742.998887 | [4] | 8 nanometer |
| 2 | navis.TreeNeuron | 754538881 | 754538881 | 4890 | None | 626 | 642 | 291434.992623 | [703] | 8 nanometer |
Exercises:
- Select the first and the last neuron in the neuronlist
- Select all neurons with a soma
- Select all neurons with a soma and less than 300,000 cable length
Further reading: https://navis.readthedocs.io/en/latest/source/tutorials/neurons_intro.html
3.6 - Plotting Neurons with Navis
Plotting
navis lets you plot neurons in 2D using matplotlib (nice for figures), and in 3D using either plotly when in a notebook environment like Deepnote or using a vispy-based 3D viewer when using a Python terminal.
import navis
# This is relevant because Deepnote does not (yet) support fancy progress bars
navis.set_pbars(jupyter=False)
# Load one of the example neurons shipped with navis
n = navis.example_neurons(1, kind='skeleton')
WARNING: Could not load OpenGL library.
# Make a 2d plot
fig, ax = navis.plot2d(n)
# Note that this is equivalent to
# fig, ax = n.plot2d()

If you have seen an olfactory projection neuron before, you might have noticed that this neuron is upside-down. That’s because hemibrain neurons have an odd orienation in that the anterior-posterior axis is not the z- but the y-axis (they were imaged from above).
For us that just means we have to turn the camera ourselves if we want a frontal view:
# Make a 2d plot
fig, ax = navis.plot2d(n)
# Change camera (azimuth + elevation)
ax.azim, ax.elev = -90, -90

Let’s do the same in 3d:
# Get a list of neurons
nl = navis.example_neurons(5)
# Plot
navis.plot3d(nl, width=1000)
Navigation:
- left click and drag to rotate (select “Orbital rotation” above the legend to make your life easier)
- mousewheel to zoom
- middle-mouse + drag to translate
- click legend items (single or double) to hide/unhide
Above plots are very basic examples but there are a ton of ways to tweak things to your liking. For a full list of parameters check out the docs for plot2d and plot3d.
Let’s for example change the colors. In general, colors can be:
- a string - e.g.
"red"or just"r" - an rgb/rgba tuple - e.g.
(1, 0, 0)for red
# Plot all neurons in red
fig, ax = navis.plot2d(n, color='r')
ax.azim, ax.elev = -90, -90

# Plot all neurons in red (color as tuple)
fig, ax = navis.plot2d(n, color=(1, 0, 0, 1))
ax.azim, ax.elev = -90, -90

When plotting multiple neurons you can either use:
- a single color (
"r"or(1, 0, 0)) -> assigned to all neurons - a list of colors (
['r', 'yellow', (0, 0, 1)]) with a color for each neuron - a dictionary mapping neuron IDs to colors (
{1734350788: 'r', 1734350908: (1, 0, 1)}) - the name of a
matplotliborseaborncolor palette
# Plot with a specific color palette
navis.plot3d(nl, color='jet')
Exercises:
- Assign rainbow colors -
"red","orange","yellow","green"and"blue"- as list - Use a dictionary to make neurons
1734350788and1734350908green, and neurons722817260,754534424and754538881red
Volumes
plot2d and plot3d also let you plot meshes. Internally these are represented as navis.Volumes (a subclass of trimesh.Trimesh):
# navis ships with a neuropil volume (in hemibrain space)
vol = navis.example_volume('neuropil')
vol
<navis.Volume(name=neuropil, color=(0.85, 0.85, 0.85, 0.2), vertices.shape=(8997, 3), faces.shape=(18000, 3))>
To plot, simply pass it to the respective plotting function:
navis.plot3d([nl, vol])
Under the hood, Volumes are treated a bit differently from neurons. So if you want to change the color, you need to do so on the object:
# Give the neuropil a reddish color
vol.color = (1, .8, .8, .4)
navis.plot3d([nl, vol], width=800)
Scatter plots
Because scatter plots are a common way of visualizing 3D data, both plot2d and plot3d provide a quick interface: (N, 3) numpy arrays and pandas.DataFrames with x, y, and z columns are interpreted as data for a scatter plot:
# Get all branch points from the node table
bp = n.branch_points
bp.head()
| node_id | label | x | y | z | radius | parent_id | type | |
|---|---|---|---|---|---|---|---|---|
| 5 | 6 | 5 | 15678.400391 | 37086.300781 | 28349.400391 | 48.011600 | 5 | branch |
| 8 | 9 | 5 | 15159.400391 | 36641.500000 | 28392.900391 | 231.296997 | 8 | branch |
| 9 | 10 | 5 | 15144.000000 | 36710.000000 | 28142.000000 | 186.977005 | 9 | branch |
| 10 | 11 | 5 | 15246.400391 | 36812.398438 | 28005.500000 | 104.261002 | 10 | branch |
| 11 | 12 | 5 | 15284.000000 | 36850.000000 | 27882.000000 | 53.245602 | 11 | branch |
# Since `bp` contains x/y/z columns, we can pass it directly to the plotting functions
navis.plot3d([n, bp],
c='k', # make the neuron black
scatter_kws=dict(color='r') # make the markers red
)
Fine-tuning figures
plot2d and plot3d provide a high-level interface to get your neurons on/in a matplotlib or a plotly figure, respectively. You can always use lower-level matplotlib/plotly interfaces directly to add more data or manipulate the figure. Just a cheap example:
# Plot neuron on a matplotlib figure
fig, ax = navis.plot2d(n, color='r')
# Show the neuron at a slight angle
ax.azim, ax.elev = -60, -60
# Zoom out a bit more
ax.dist = 8 # default is 7
# Unhide axes
ax.set_axis_on()
# Label the axes
ax.set_xlabel('x-axis [8 nm voxels]')
ax.set_ylabel('y-axis [8 nm voxels]')
ax.set_zlabel('z-axis [8 nm voxels]')
Text(0.5, 0, 'z-axis [8 nm voxels]')
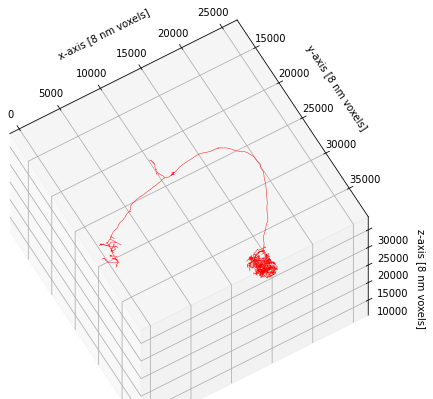
This concludes this brief introduction to plotting but just to note that plot2d and plot3d have a lot of additional functionality to customize the way neurons are plotted. If you have time on your hands, I recommend you check out and play around with the available parameters (e.g. linewidth, color_by, shade_by, linestyle).
3.7 - pymaid
Overview
pymaid lets you interface with a CATMAID server. It’s built on top of navis and returns data (neurons, volumes) in a way that you can plug them straight into navis to use features such as plotting.
Official documentation here.
Connecting
The VFB CATMAID servers (see here for what’s available) are public and don’t require an API token for read-only access which makes connecting simple:
import pymaid
import navis
navis.set_pbars(jupyter=False)
pymaid.set_pbars(jupyter=False)
# Connect to the VFB CATMAID server hosting the FAFB data
rm = pymaid.connect_catmaid(server="https://fafb.catmaid.virtualflybrain.org/", api_token=None, max_threads=10)
# Test call to see if connection works
print(f'Server is running CATMAID version {rm.catmaid_version}')
WARNING: Could not load OpenGL library.
INFO : Global CATMAID instance set. Caching is ON. (pymaid)
Server is running CATMAID version 2020.02.15-905-g93a969b37
Retrieving neurons
Let’s start with pulling a neuron based on its ID:
# Find a neuron from its ID (16) -> this is an olfactory projection neuron
n = pymaid.get_neurons(16)
n
| type | CatmaidNeuron |
|---|---|
| name | Uniglomerular mALT VA6 adPN 017 DB |
| id | 16 |
| n_nodes | 16840 |
| n_connectors | 2158 |
| n_branches | 1172 |
| n_leafs | 1230 |
| cable_length | 4003103.232861 |
| soma | [2941309] |
| units | 1 nanometer |
This neuron’s type is pymaid.CatmaidNeuron, which is a subclass of navis.TreeNeuron. The list version is pymaid.CatmaidNeuronList, which a subclass of navis.NeuronList. This adds a bit of extra functionality (such as lazy loading of data) and allows CatmaidNeuron and CatmaidNeuronList work as drop in replacements for their parent classes.
# Plot CatmaidNeuron with navis
navis.plot3d(n, width=1000, connectors=True, c='k')
get_neurons() returns neurons including their “connectors” - i.e. pre- (red) and postsynapses (blue). For this particular neuron, the published data comprehensively labels the axonal synapses but not the dendrites. Analogous to the nodes table, you can access the connectors like so:
n.connectors.head()
| node_id | connector_id | type | x | y | z | |
|---|---|---|---|---|---|---|
| 0 | 97891 | 97895 | 0 | 436882.09375 | 161840.453125 | 212160.0 |
| 1 | 2591 | 97954 | 0 | 437120.00000 | 160998.000000 | 211920.0 |
| 2 | 2665 | 98300 | 0 | 437183.75000 | 162323.515625 | 214880.0 |
| 3 | 2646 | 98373 | 0 | 437041.68750 | 162451.937500 | 214120.0 |
| 4 | 2654 | 98415 | 0 | 436760.90625 | 163689.796875 | 214440.0 |
Let’s run a bigger example and pull all data published with Bates, Schlegel et al. 2020. For this, we will use “annotations”. These are effectively text labels that group neurons together, in this case by paper. Instead of get_neurons we can use find_neurons to avoid downloading unnecessary data.
bates = pymaid.find_neurons(annotations='Paper: Bates and Schlegel et al 2020')
len(bates)
INFO : Found 583 neurons matching the search parameters (pymaid)
583
bates is a CatmaidNeuronList containing 583 neurons. Importantly pymaid has not yet loaded any data other than names! Note all the “NAs” in the summary:
bates.head()
| type | name | skeleton_id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57316 2863105 ML | 2863104 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 1 | CatmaidNeuron | Uniglomerular mALT DA3 adPN 57350 HG | 57349 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 2 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57354 GA | 57353 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 3 | CatmaidNeuron | Uniglomerular mALT VA6 adPN 017 DB | 16 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 4 | CatmaidNeuron | Uniglomerular mALT VA5 lPN 57362 ML | 57361 | NA | NA | NA | NA | NA | NA | 1 nanometer |
We could have used pymaid.get_neurons(annotations='Paper: Bates and Schlegel et al 2020') instead to load all data up-front, but this would increase memory usage.
The CatmaidNeuronList we have created will lazy load data from the server when required.
# Access the first neuron's nodes
# -> this will trigger a data download
_ = bates[0].nodes
# Run summary again
bates.head()
| type | name | skeleton_id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57316 2863105 ML | 2863104 | 6774 | 470 | 280 | 292 | 1522064.513255 | [3245741] | 1 nanometer |
| 1 | CatmaidNeuron | Uniglomerular mALT DA3 adPN 57350 HG | 57349 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 2 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57354 GA | 57353 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 3 | CatmaidNeuron | Uniglomerular mALT VA6 adPN 017 DB | 16 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 4 | CatmaidNeuron | Uniglomerular mALT VA5 lPN 57362 ML | 57361 | NA | NA | NA | NA | NA | NA | 1 nanometer |
We have now loaded data for the first neuron.
Next we willl find and plot all uniglomelar DA1 projection neurons by their name.
# Name will be match pattern "Uniglomerular {tract} DA1 {lineage}"
import re
prog = re.compile("Uniglomerular(.*?) DA1 ")
# Match all neuron names in `bates` against that pattern
is_da1 = list(map(lambda x: prog.match(x) != None, bates.name))
# Subset list
da1 = bates[is_da1]
da1.head()
| type | name | skeleton_id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57316 2863105 ML | 2863104 | 6774 | 470 | 280 | 292 | 1522064.513255 | [3245741] | 1 nanometer |
| 1 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57354 GA | 57353 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 2 | CatmaidNeuron | Uniglomerular mALT DA1 lPN 57382 ML | 57381 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 3 | CatmaidNeuron | Uniglomerular mlALT DA1 vPN mlALTed Milk 23348... | 2334841 | NA | NA | NA | NA | NA | NA | 1 nanometer |
| 4 | CatmaidNeuron | Uniglomerular mALT DA1 lPN PN021 2345090 DB RJVR | 2345089 | NA | NA | NA | NA | NA | NA | 1 nanometer |
# Plot neurons by their lineage
for n in da1:
# Split name into components and keep the lineage
n.lineage = n.name.split(' ')[3]
# Generate a color per lineage
import seaborn as sns
import numpy as np
lineages = np.unique(da1.lineage)
lin_cmap = dict(zip(lineages, sns.color_palette('muted', len(lineages))))
neuron_cmap = {n.id: lin_cmap[n.lineage] for n in da1}
navis.plot3d(da1, color=neuron_cmap, hover_name=True)
Let’s add the neuropil meshes. These are called “volumes” on the CATMAID servers. To find out what’s available:
vols = pymaid.get_volume()
vols.head()
INFO : Retrieving list of available volumes. (pymaid)
| id | name | comment | user_id | editor_id | project_id | creation_time | edition_time | annotations | area | volume | watertight | meta_computed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 439 | v14.neuropil | None | 55 | 247 | 1 | 2017-10-05T21:01:18.683Z | 2018-08-30T17:21:20.910Z | None | 6.377313e+11 | 1.533375e+16 | False | True |
| 1 | 440 | AME_R | Accessory medulla right | 55 | 55 | 1 | 2017-10-08T13:54:03.279Z | 2017-10-08T13:54:03.279Z | None | 1.894095e+09 | 4.799292e+12 | True | True |
| 2 | 441 | LO_R | Lobula right | 55 | 55 | 1 | 2017-10-08T13:54:03.840Z | 2017-10-08T13:54:03.840Z | None | 4.103282e+10 | 5.790708e+14 | True | True |
| 3 | 442 | NO | Noduli | 55 | 55 | 1 | 2017-10-08T13:54:04.084Z | 2017-10-08T13:54:04.084Z | None | 3.955158e+09 | 1.796395e+13 | True | True |
| 4 | 443 | BU_R | Bulb right | 55 | 55 | 1 | 2017-10-08T13:54:04.263Z | 2017-10-08T13:54:04.263Z | None | 1.445868e+09 | 4.109262e+12 | True | True |
# Get the neuropil volume
v14neuropil = pymaid.get_volume('v14.neuropil')
# Make it slightly more transparent
v14neuropil.color = (.8, .8, .8, .3)
INFO : Cached data used. Use `pymaid.clear_cache()` to clear. (pymaid)
# Plot with neuropil volume
navis.plot3d([da1, v14neuropil], color=neuron_cmap)
Suggested exercises:
- find all uniglomerular projection neurons (name starts with
Uniglomerular) - calculate the number of pre-/post-synapses in the right lateral horn (LH) (use
pymaid.get_volumeandnavis.in_volume) - group the neurons by glomerulus based on label (nomenclature is
Uniglomerular {tract} {glomerulus} {lineage} {metadata}) - plot LH pre- vs post-synapses in a scatter plot (e.g. using
seaborn.scatterplot)
Pulling connectivity
CATMAID lets you fetch connectivity data either as a list of up- and downstream partners or as whole adjacency matrices.
# Pull downstream partners of DA1 PNs
da1_ds = pymaid.get_partners(da1,
threshold=3, # anything with >= 3 synapses
directions=['outgoing'] # downstream partners only
)
# Result is a pandas DataFrame
da1_ds.head()
INFO : Fetching connectivity table for 17 neurons (pymaid)
INFO : Done. Found 0 pre-, 270 postsynaptic and 0 gap junction-connected neurons (pymaid)
| neuron_name | skeleton_id | num_nodes | relation | 2863104 | 57353 | 57381 | 2334841 | 2345089 | 27295 | ... | 2319457 | 4207871 | 755022 | 2379517 | 61221 | 3239781 | 2381753 | 57311 | 57323 | total | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | Uniglomerular mlALT DA1 vPN mlALTed Milk 18114... | 1811442 | 11769 | downstream | 30 | 3 | 4 | 0 | 0 | 15 | ... | 0 | 0 | 32 | 0 | 26 | 0 | 0 | 21 | 20 | 151.0 |
| 1 | Uniglomerular mlALT DA1 vPN mlALTed Milk 23348... | 2334841 | 6362 | downstream | 0 | 0 | 0 | 0 | 14 | 0 | ... | 22 | 17 | 0 | 28 | 0 | 26 | 32 | 0 | 0 | 139.0 |
| 2 | LHAV4a4#1 1911125 FML PS RJVR | 1911124 | 6969 | downstream | 23 | 6 | 9 | 0 | 0 | 5 | ... | 0 | 0 | 19 | 0 | 13 | 0 | 0 | 19 | 15 | 109.0 |
| 3 | LHAV2a3#1 1870231 RJVR AJES PS | 1870230 | 14820 | downstream | 5 | 23 | 28 | 0 | 0 | 10 | ... | 0 | 0 | 19 | 0 | 7 | 0 | 0 | 5 | 7 | 105.0 |
| 4 | LHAV4c1#1 488056 downstream DA1 GSXEJ | 488055 | 12137 | downstream | 15 | 3 | 0 | 0 | 0 | 16 | ... | 0 | 0 | 15 | 0 | 15 | 0 | 0 | 17 | 11 | 92.0 |
5 rows × 22 columns
Each row is a synaptic downstream partner of our query DA1 neurons. The columns to the left contain the synapses they receive from individual query neurons. For example 1811442 (first row) receives 30 synapses from the DA1 PN with ID 2863104.
# Get an adjacency matrix between all Bates, Schlegel et al. neurons
adj = pymaid.adjacency_matrix(bates)
adj.head()
| targets | 2863104 | 57349 | 57353 | 16 | 57361 | 15738898 | 57365 | 4182038 | 3813399 | 11524119 | ... | 57323 | 4624362 | 1853423 | 2842610 | 57333 | 4624374 | 3080183 | 57337 | 4624378 | 57341 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| sources | |||||||||||||||||||||
| 2863104 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 2.0 | 0.0 | 12.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 57349 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 57353 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 5.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 16 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 57361 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
5 rows × 583 columns
# Plot a quick & dirty adjacency matrix
import seaborn as sns
ax = sns.clustermap(adj, vmax=10, cmap='Greys')
/shared-libs/python3.7/py/lib/python3.7/site-packages/seaborn/matrix.py:649: UserWarning:
Clustering large matrix with scipy. Installing `fastcluster` may give better performance.

We can also ask for where in space specific connections are made:
# Axo-axonic connections between two different types of DA1 PNs
cn = pymaid.get_connectors_between(2863104, 1811442)
cn.head()
| connector_id | connector_loc | node1_id | source_neuron | confidence1 | creator1 | node1_loc | node2_id | target_neuron | confidence2 | creator2 | node2_loc | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 6736296 | [359448.44, 159319.03, 150560.0] | 3163408 | 2863104 | 5 | NaN | [359487.3, 159145.66, 150600.0] | 6736298 | 1811442 | 5 | NaN | [359611.9, 159541.48, 150560.0] |
| 1 | 6795172 | [356041.88, 149555.53, 147920.0] | 6795195 | 2863104 | 5 | NaN | [354724.44, 149284.1, 147920.0] | 6795153 | 1811442 | 5 | NaN | [356366.16, 149854.86, 147920.0] |
| 2 | 6795291 | [355189.5, 150232.48, 148240.0] | 6795293 | 2863104 | 5 | NaN | [354595.62, 149464.8, 148240.0] | 6795214 | 1811442 | 5 | NaN | [355472.28, 150294.75, 148160.0] |
| 3 | 6795747 | [355030.4, 154047.86, 145800.0] | 6795749 | 2863104 | 5 | NaN | [355045.38, 154180.1, 145800.0] | 6795745 | 1811442 | 5 | NaN | [355024.44, 153945.73, 145760.0] |
| 4 | 6797452 | [353221.4, 148570.9, 147320.0] | 6797456 | 2863104 | 5 | NaN | [354213.9, 148397.44, 147320.0] | 6797437 | 1811442 | 5 | NaN | [353447.6, 148704.88, 147560.0] |
# Visualize
points = np.vstack(cn.connector_loc)
navis.plot3d([da1.idx[[2863104, 1811442]], # plot the two neurons
points], # plot the points of synaptic contacts as scatter
scatter_kws=dict(name="synaptic contacts")
)
3.8 - NBLAST
Overview
NBLAST (Costa et al., 2016) is a method to quantify morphological similarity. It works on “dotprops” which represent neurons as tangent vectors. For each tangent vector in the query neuron, NBLAST finds the closest tangent vector in the target neuron and calculates a score from the distance between and the dotproduct of the two vectors. The final NBLAST score is the sum over all query-target vector pairs. Typically, this score is normalized to a self-self comparison (i.e. a perfect match would be 1).

Finding Matching Neurons
VFB computes NBLAST scores for all neurons in its database. So if all you want is a list of similar neurons, it’s fastest (and easiest) to get those directly from VFB.
# Import libs and initialise API objects
from vfb_connect.cross_server_tools import VfbConnect
import navis.interfaces.neuprint as neu
import pandas as pd
import navis
navis.set_pbars(jupyter=False)
vc = VfbConnect()
client = neu.Client('https://neuprint.janelia.org', dataset='hemibrain:v1.1')
First let’s write a function to grab VFB neurons and turn them into navis neurons. This is modified from vc.neo_query_wrapper.get_images.
import requests
from tqdm import tqdm
def get_vfb_neurons(vfb_ids, template='JRC2018Unisex'):
"""Load neurons from VFB as navis.TreeNeurons."""
vfb_ids = list(navis.utils.make_iterable(vfb_ids))
inds = vc.neo_query_wrapper.get_anatomical_individual_TermInfo(short_forms=vfb_ids)
nl = []
# Note: queries should be parallelized in the future
# -> for pymaid I use request-futures which works really well
for i in tqdm(inds, desc='Loading VFB neurons', leave=False):
if not ('has_image' in i['term']['core']['types']):
continue
label = i['term']['core']['label']
image_matches = [x['image'] for x in i['channel_image']]
if not image_matches:
continue
for imv in image_matches:
if imv['template_anatomy']['label'] == template:
r = requests.get(imv['image_folder'] + '/volume.swc')
### Slightly dodgy warning - could mask network errors
if not r.ok:
warnings.warn("No '%s' file found for '%s'." % (image_type, label))
continue
# `id` should ideally be unique but that's not enforced
n = navis.TreeNeuron(r.text, name=label, id=i['term']['core']['short_form'])
# This registers attributes that you want to show in the summary
# alternatively we could have a generic attribute like so:
# n.template = template
n._register_attr("template", template, temporary=False)
# I assume all available templates are in microns?
# @Robbie -do we have this metadata on templates? If not, we should add ASAP.
n.units = '1 micron'
nl.append(n)
return navis.NeuronList(nl)
# Search for similar neurons using the VFB ID of an ellipsoid body neuron in FAFB
# We got that ID from a search on the VFB website
similar_to_EPG6L1 = vc.get_similar_neurons('VFB_001012ay')
similar_to_EPG6L1
| id | NBLAST_score | label | types | source_id | accession_in_source | |
|---|---|---|---|---|---|---|
| 0 | VFB_jrchjtk6 | 0.525 | EPG(PB08)_L6 - 541870397 | [EB-PB 1 glomerulus-D/Vgall neuron] | neuprint_JRC_Hemibrain_1point1 | 541870397 |
| 1 | VFB_jrchjtk8 | 0.524 | EPG(PB08)_L6 - 912601268 | [EB-PB 1 glomerulus-D/Vgall neuron] | neuprint_JRC_Hemibrain_1point1 | 912601268 |
| 2 | VFB_jrchjtkb | 0.504 | EPG(PB08)_L6 - 788794171 | [EB-PB 1 glomerulus-D/Vgall neuron] | neuprint_JRC_Hemibrain_1point1 | 788794171 |
| 3 | VFB_jrchjtji | 0.481 | EL(EQ5)_L - 1036753721 | [EBw.AMP.s-Dga-s.b neuron] | neuprint_JRC_Hemibrain_1point1 | 1036753721 |
# Read those neurons from VFB
query = get_vfb_neurons('VFB_001012ay')
matches = get_vfb_neurons(similar_to_EPG6L1.id.values)
matches
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | template | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | EPG(PB08)_L6 - 912601268 | VFB_jrchjtk8 | 9819 | None | 1000 | 1015 | 2211.292188 | None | 1 micron | JRC2018Unisex |
| 1 | navis.TreeNeuron | EL(EQ5)_L - 1036753721 | VFB_jrchjtji | 13864 | None | 1942 | 1974 | 3403.311508 | [1] | 1 micron | JRC2018Unisex |
| 2 | navis.TreeNeuron | EPG(PB08)_L6 - 788794171 | VFB_jrchjtkb | 9651 | None | 962 | 987 | 2125.364857 | None | 1 micron | JRC2018Unisex |
| 3 | navis.TreeNeuron | EPG(PB08)_L6 - 541870397 | VFB_jrchjtk6 | 11042 | None | 1099 | 1127 | 2432.101351 | [1] | 1 micron | JRC2018Unisex |
# Define colors such that query is black
import seaborn as sns
colors = ['k'] + sns.color_palette('muted', len(matches))
navis.plot3d([query, matches], color=colors)
These like a good match!
Clustering Groups of Neurons
Now imagine you have a set of neurons and you want to group them into morphologically similar “types”. To demonstrate this, we will take some ellipsoid body neurons from VFB and cluster them using NBLAST.
# Find instances of EB-PB-gall neurons
eb_pb_ids = pd.DataFrame.from_records(vc.get_instances("adult ellipsoid body-protocerebral bridge-gall neuron"))
eb_pb_ids.head()
| label | symbol | id | tags | parents_label | parents_id | data_source | accession | templates | dataset | license | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | EPGt(PB09)_L9 - 1168664447 | VFB_jrchjtlc | Entity|has_image|Adult|Anatomy|has_neuron_conn... | EB slice 8-GA-PB slice 9 neuron | FBbt_00111423 | neuprint_JRC_Hemibrain_1point1 | 1168664447 | JRC_FlyEM_Hemibrain|JRC2018Unisex | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... | |
| 1 | EPGt(PB09)_R9 - 1219069439 | VFB_jrchjtl9 | Entity|has_image|Adult|Anatomy|has_neuron_conn... | EB slice 8-GA-PB slice 9 neuron | FBbt_00111423 | neuprint_JRC_Hemibrain_1point1 | 1219069439 | JRC2018Unisex|JRC_FlyEM_Hemibrain | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... | |
| 2 | EPG(PB08)_L2 - 697001770 | VFB_jrchjtkw | Entity|has_image|Adult|Anatomy|has_neuron_conn... | EB-PB 1 glomerulus-D/Vgall neuron | FBbt_00047030 | neuprint_JRC_Hemibrain_1point1 | 697001770 | JRC2018Unisex|JRC_FlyEM_Hemibrain | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... | |
| 3 | EPG(PB08)_R8 - 1125964814 | VFB_jrchjtl5 | Entity|has_image|Adult|Anatomy|has_neuron_conn... | EB-PB 1 glomerulus-D/Vgall neuron | FBbt_00047030 | neuprint_JRC_Hemibrain_1point1 | 1125964814 | JRC2018Unisex|JRC_FlyEM_Hemibrain | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... | |
| 4 | EPG(PB08)_L6 - 912601268 | VFB_jrchjtk8 | Entity|has_image|Adult|Anatomy|has_neuron_conn... | EB-PB 1 glomerulus-D/Vgall neuron | FBbt_00047030 | neuprint_JRC_Hemibrain_1point1 | 912601268 | JRC2018Unisex|JRC_FlyEM_Hemibrain | Xu2020NeuronsV1point1 | https://creativecommons.org/licenses/by/4.0/le... |
# Get the neurons from VFB
eb_pb = get_vfb_neurons(eb_pb_ids.id.values)
eb_pb
Loading VFB neurons: 100%|██████████| 62/62 [01:04<00:00, 1.04s/it]
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | EPG(PB08)_R5 - 725951521 | VFB_jrchjtk5 | 11097 | None | 1223 | 1251 | 2221.279929 | [1] | 1 dimensionless |
| 1 | navis.TreeNeuron | EPG(PB08)_R2 - 632544268 | VFB_jrchjtkt | 12861 | None | 1475 | 1507 | 2664.323311 | [1] | 1 dimensionless |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 60 | navis.TreeNeuron | EPG(PB08)_R5 - 694920753 | VFB_jrchjtk7 | 11358 | None | 1348 | 1376 | 2335.616245 | [1] | 1 dimensionless |
| 61 | navis.TreeNeuron | EPG(PB08)_R6 - 910438331 | VFB_jrchjtkc | 10983 | None | 1086 | 1107 | 2340.485198 | [1] | 1 dimensionless |
Now that we have the skeletons, we need to turn them into dotprops. Keep in mind that:
- NBLAST is optimized for data in microns. Since our neurons are in the
JRC2018Unisextemplate, they already are. - Neurons should have the same sampling rate (i.e. nodes per micron). To ensure this we will resample the neurons to 1 point per micron.
- We won’t be doing this this time around but we typically simplify the neurons slightly by e.g. pruning small twigs (
navis.prune_twigs()) to reduce noise.
eb_dps = navis.make_dotprops(eb_pb, resample=1, k=5)
# Note this NeuronList /contains dotprops now
eb_dps
| type | name | id | k | units | n_points | |
|---|---|---|---|---|---|---|
| 0 | navis.Dotprops | EPG(PB08)_R5 - 725951521 | VFB_jrchjtk5 | 5 | 1 dimensionless | 2683 |
| 1 | navis.Dotprops | EPG(PB08)_R2 - 632544268 | VFB_jrchjtkt | 5 | 1 dimensionless | 3152 |
| ... | ... | ... | ... | ... | ... | ... |
| 60 | navis.Dotprops | EPG(PB08)_R5 - 694920753 | VFB_jrchjtk7 | 5 | 1 dimensionless | 2924 |
| 61 | navis.Dotprops | EPG(PB08)_R6 - 910438331 | VFB_jrchjtkc | 5 | 1 dimensionless | 2486 |
To illustrate what dotprops are, lets visualize one side-by-side with its skeleton.
sk = eb_pb[0].copy()
dp = eb_dps[0].copy()
# Slightly offset the dotprops by a micron
dp.points[:, 0] += 1
navis.plot3d([sk, dp], color=['k', 'r'])
Now we can run the NBLAST. Note that in this case, we run an all-by-all NBLAST which means that we don’t need to worry about the directionality - i.e. whether we NBLAST A->B or B->A.
scores = navis.nblast_allbyall(eb_dps)
scores.head()
| VFB_jrchjtk5 | VFB_jrchjtkt | VFB_001012ba | VFB_jrchjtkj | VFB_jrchjtkp | VFB_jrchjtke | VFB_jrchjtkr | VFB_jrchjtkb | VFB_00004319 | VFB_00009714 | ... | VFB_jrchjtk8 | VFB_001012bp | VFB_jrchjtk9 | VFB_jrchjtki | VFB_jrchjtl3 | VFB_jrchjtkv | VFB_001012b9 | VFB_001012bb | VFB_jrchjtk7 | VFB_jrchjtkc | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| VFB_jrchjtk5 | 1.000000 | -0.083960 | 0.427659 | 0.058210 | 0.021904 | 0.484210 | 0.168235 | 0.178954 | 0.065457 | 0.355732 | ... | 0.130788 | 0.528314 | 0.569923 | -0.029195 | -0.130578 | -0.072185 | 0.381668 | 0.139970 | 0.816455 | 0.504740 |
| VFB_jrchjtkt | -0.062465 | 1.000000 | -0.030980 | -0.124326 | 0.507521 | -0.122518 | -0.143298 | 0.091243 | -0.124431 | -0.142560 | ... | 0.172337 | -0.059201 | 0.078412 | -0.116040 | 0.129102 | -0.034670 | -0.053382 | 0.103465 | -0.063922 | -0.073508 |
| VFB_001012ba | 0.129771 | -0.115879 | 1.000000 | 0.374231 | -0.091531 | 0.549519 | 0.450368 | -0.130200 | 0.161574 | 0.112378 | ... | -0.127990 | 0.786001 | 0.013910 | 0.330330 | -0.000875 | 0.212914 | 0.817302 | -0.110084 | 0.200497 | 0.569953 |
| VFB_jrchjtkj | -0.027448 | -0.219602 | 0.483341 | 1.000000 | -0.031111 | 0.443643 | 0.461981 | -0.025682 | 0.061803 | 0.434210 | ... | -0.034895 | 0.389407 | -0.091165 | 0.819716 | 0.217721 | 0.552982 | 0.447278 | -0.053055 | 0.045722 | 0.485596 |
| VFB_jrchjtkp | -0.158602 | 0.493915 | -0.161524 | -0.103061 | 1.000000 | -0.212700 | -0.221368 | 0.405601 | -0.163415 | -0.130491 | ... | 0.515069 | -0.170831 | 0.024440 | -0.095100 | -0.032720 | -0.082523 | -0.182386 | 0.412998 | -0.158171 | -0.197676 |
5 rows × 62 columns
Next, we need to take those forward scores (A->B) and symmetrize by calculating mean scores (A<->B).
# Symmetrize scores
scores_mean = (scores + scores.T) / 2
scores_mean.head()
| VFB_jrchjtk5 | VFB_jrchjtkt | VFB_001012ba | VFB_jrchjtkj | VFB_jrchjtkp | VFB_jrchjtke | VFB_jrchjtkr | VFB_jrchjtkb | VFB_00004319 | VFB_00009714 | ... | VFB_jrchjtk8 | VFB_001012bp | VFB_jrchjtk9 | VFB_jrchjtki | VFB_jrchjtl3 | VFB_jrchjtkv | VFB_001012b9 | VFB_001012bb | VFB_jrchjtk7 | VFB_jrchjtkc | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| VFB_jrchjtk5 | 1.000000 | -0.073212 | 0.278715 | 0.015381 | -0.068349 | 0.411257 | 0.098761 | 0.142277 | 0.032064 | 0.187264 | ... | 0.070828 | 0.437337 | 0.572156 | -0.043306 | -0.140155 | -0.092071 | 0.271765 | 0.131925 | 0.820455 | 0.398975 |
| VFB_jrchjtkt | -0.073212 | 1.000000 | -0.073429 | -0.171964 | 0.500718 | -0.142672 | -0.164992 | 0.062182 | -0.115399 | -0.162311 | ... | 0.176281 | -0.099006 | 0.074666 | -0.162002 | 0.097262 | -0.082222 | -0.089534 | 0.120931 | -0.075555 | -0.114645 |
| VFB_001012ba | 0.278715 | -0.073429 | 1.000000 | 0.428786 | -0.126527 | 0.667980 | 0.527915 | -0.128599 | 0.220682 | 0.068423 | ... | -0.143015 | 0.792216 | 0.070675 | 0.389590 | 0.018090 | 0.176492 | 0.826461 | -0.102122 | 0.332148 | 0.684591 |
| VFB_jrchjtkj | 0.015381 | -0.171964 | 0.428786 | 1.000000 | -0.067086 | 0.436914 | 0.494708 | -0.031511 | -0.030539 | 0.489038 | ... | -0.045253 | 0.396773 | -0.069074 | 0.824681 | 0.168068 | 0.508538 | 0.395092 | -0.062799 | 0.100087 | 0.509110 |
| VFB_jrchjtkp | -0.068349 | 0.500718 | -0.126527 | -0.067086 | 1.000000 | -0.164867 | -0.203583 | 0.413305 | -0.171488 | -0.124160 | ... | 0.537125 | -0.109346 | 0.020020 | -0.074199 | -0.011358 | -0.093459 | -0.133473 | 0.435029 | -0.062292 | -0.161714 |
5 rows × 62 columns
From here on out, we use cookie cutter scipy to produce a hierarchical clustering. First we need to convert those similarity scores to distances because that’s what scipy operates on:
dist_mean = (scores_mean - 1) * -1
dist_mean.head()
| VFB_jrchjtk5 | VFB_jrchjtkt | VFB_001012ba | VFB_jrchjtkj | VFB_jrchjtkp | VFB_jrchjtke | VFB_jrchjtkr | VFB_jrchjtkb | VFB_00004319 | VFB_00009714 | ... | VFB_jrchjtk8 | VFB_001012bp | VFB_jrchjtk9 | VFB_jrchjtki | VFB_jrchjtl3 | VFB_jrchjtkv | VFB_001012b9 | VFB_001012bb | VFB_jrchjtk7 | VFB_jrchjtkc | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| VFB_jrchjtk5 | -0.000000 | 1.073212 | 0.721285 | 0.984619 | 1.068349 | 0.588743 | 0.901239 | 0.857723 | 0.967936 | 0.812736 | ... | 0.929172 | 0.562663 | 0.427844 | 1.043306 | 1.140155 | 1.092071 | 0.728235 | 0.868075 | 0.179545 | 0.601025 |
| VFB_jrchjtkt | 1.073212 | -0.000000 | 1.073429 | 1.171964 | 0.499282 | 1.142672 | 1.164992 | 0.937818 | 1.115399 | 1.162311 | ... | 0.823719 | 1.099006 | 0.925334 | 1.162002 | 0.902738 | 1.082222 | 1.089534 | 0.879069 | 1.075555 | 1.114645 |
| VFB_001012ba | 0.721285 | 1.073429 | -0.000000 | 0.571214 | 1.126527 | 0.332020 | 0.472085 | 1.128599 | 0.779318 | 0.931577 | ... | 1.143015 | 0.207784 | 0.929325 | 0.610410 | 0.981910 | 0.823508 | 0.173539 | 1.102122 | 0.667852 | 0.315409 |
| VFB_jrchjtkj | 0.984619 | 1.171964 | 0.571214 | -0.000000 | 1.067086 | 0.563086 | 0.505292 | 1.031511 | 1.030539 | 0.510962 | ... | 1.045253 | 0.603227 | 1.069074 | 0.175319 | 0.831932 | 0.491462 | 0.604908 | 1.062799 | 0.899913 | 0.490890 |
| VFB_jrchjtkp | 1.068349 | 0.499282 | 1.126527 | 1.067086 | -0.000000 | 1.164867 | 1.203583 | 0.586695 | 1.171488 | 1.124160 | ... | 0.462875 | 1.109346 | 0.979980 | 1.074199 | 1.011358 | 1.093459 | 1.133473 | 0.564971 | 1.062292 | 1.161714 |
5 rows × 62 columns
from scipy.spatial.distance import squareform
from scipy.cluster.hierarchy import linkage, dendrogram, fcluster
# This turns the square distance matrix into 1-d vector form
dist_sq = squareform(dist_mean)
# This does the actual hierarchical clustering
Z = linkage(dist_sq, method='ward', optimal_ordering=True)
# Plot a quick dendrogram
import matplotlib.pyplot as plt
import seaborn as sns
# Generate the canvas
fig, ax = plt.subplots(figsize=(12, 5))
# Make some more meaningful labels
vfb2name = dict(zip(eb_pb.id, eb_pb.name))
labels = dist_mean.index.map(lambda x: vfb2name[x].split(' - ')[0])
# Plot the dendrogram
dn = dendrogram(Z, ax=ax, labels=labels, color_threshold=.65, above_threshold_color='lightgrey')
# Make xticks biggger
ax.set_xticklabels(ax.get_xticklabels(), size=11)
# Clean axes
sns.despine(bottom=True)
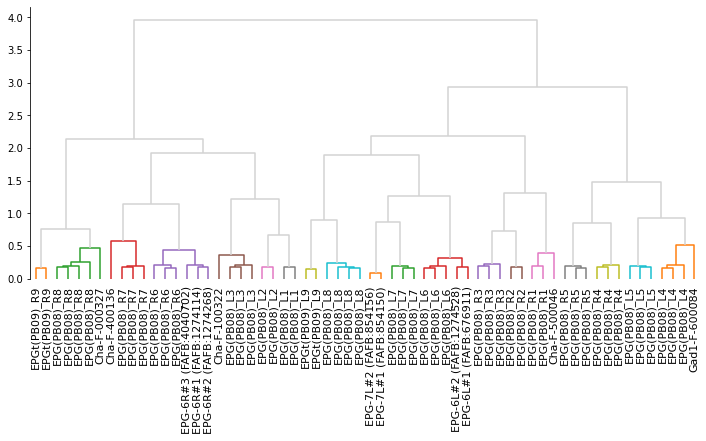
# Make cluster matching the color threshold in the dendrogram
clusters = fcluster(Z, t=.65, criterion='distance')
clusters
array([16, 14, 4, 5, 11, 4, 3, 12, 2, 19, 13, 4, 8, 3, 19, 2, 5,
13, 3, 11, 17, 14, 16, 6, 12, 15, 18, 11, 12, 15, 15, 18, 5, 10,
9, 2, 19, 8, 1, 9, 19, 3, 1, 10, 2, 9, 7, 18, 9, 2, 17,
13, 12, 4, 17, 5, 7, 6, 4, 12, 16, 4], dtype=int32)
# We have to downsample the neurons for visualization
# because DeepNote enforces a limit on the size of plots
eb_pb_ds = eb_pb.downsample(10)
# Make a color per cluster
palette = sns.color_palette('tab20', max(clusters))
cmap = dict(zip(range(1, max(clusters) + 1), palette))
colors = [cmap[i] for i in clusters]
navis.plot3d(eb_pb_ds, color=colors)