neuprint
hemibrain data
The Janelia hemibrain dataset (Scheffer et al., 2020) is accessible via neuprint at https://neuprint.janelia.org. The webinterface lets you run a couple pre-built queries but you can also run custom queries directly against the underlying neo4j graph data base. It’s worth looking at their data model and reading up on how neo4j “cyphers” (i.e. queries) work.
To programmatically interface with neuprint, we will use neuprint-python (link). It requires an API token which you can get via the website and is bound to the Google account that you use to log into neuprint. For this workshop we provide such a token as environment variable but you will need to start using your own token after the workshop is over.
neuprint-python
First we have to initialize the connection.
import neuprint as neu
client = neu.Client('https://neuprint.janelia.org', dataset='hemibrain:v1.1',
token="eyJhbGciOiJIUzI1NiIsInR5cCI6IkpXVCJ9.eyJlbWFpbCI6InZmYndvcmtzaG9wLm5ldXJvZmx5MjAyMEBnbWFpbC5jb20iLCJsZXZlbCI6Im5vYXV0aCIsImltYWdlLXVybCI6Imh0dHBzOi8vbGg2Lmdvb2dsZXVzZXJjb250ZW50LmNvbS8tWXFDN21NRXd3TlEvQUFBQUFBQUFBQUkvQUFBQUFBQUFBQUEvQU1adXVjbU5zaXhXZDRhM0VyTTQ0ODBMa2IzNDdvUlpfUS9zOTYtYy9waG90by5qcGc_c3o9NTA_c3o9NTAiLCJleHAiOjE3OTQwOTE4ODd9.ceg4mrj2o-aOhK0NHNGmBacg8R34PBPoLBwhCo4uOCQ")
Most functions in neuprint-python accept neu.NeuronCriteria which is effectively a filter for body IDs, types, etc:
help(neu.NeuronCriteria)
Help on class NeuronCriteria in module neuprint.neuroncriteria:
class NeuronCriteria(builtins.object)
| NeuronCriteria(matchvar='n', *, bodyId=None, instance=None, type=None, regex=False, cellBodyFiber=None, status=None, cropped=None, min_pre=0, min_post=0, rois=None, inputRois=None, outputRois=None, min_roi_inputs=1, min_roi_outputs=1, label=None, roi_req='all', client=None)
|
| Specifies which fields to filter by when searching for a Neuron (or Segment).
| This class does not send queries itself, but you use it to specify search
| criteria for various query functions.
|
| Note:
| For simple queries involving only particular bodyId(s) or type(s)/instance(s),
| you can usually just pass the ``bodyId`` or ``type`` to the query function,
| without constructing a full ``NeuronCriteria``.
|
| .. code-block:: python
|
| from neuprint import fetch_neurons, NeuronCriteria as NC
|
| # Equivalent
| neuron_df, conn_df = fetch_neurons(NC(bodyId=329566174))
| neuron_df, conn_df = fetch_neurons(329566174)
|
| # Equivalent
| # (Criteria is satisfied if either type or instance matches.)
| neuron_df, conn_df = fetch_neurons(NC(type="OA-VPM3", instance="OA-VPM3"))
| neuron_df, conn_df = fetch_neurons("OA-VPM3")
|
| Methods defined here:
|
| __eq__(self, value)
| Implement comparison between criteria.
| Note: 'matchvar' is not considered during the comparison.
|
| __init__(self, matchvar='n', *, bodyId=None, instance=None, type=None, regex=False, cellBodyFiber=None, status=None, cropped=None, min_pre=0, min_post=0, rois=None, inputRois=None, outputRois=None, min_roi_inputs=1, min_roi_outputs=1, label=None, roi_req='all', client=None)
| Except for ``matchvar``, all parameters must be passed as keyword arguments.
|
| .. note::
|
| **Options for specifying ROI criteria**
|
| The ``rois`` argument merely matches neurons that intersect the given ROIs at all
| (without distinguishing between inputs and outputs).
|
| The ``inputRois`` and ``outputRois`` arguments allow you to put requirements
| on whether or not neurons have inputs or outputs in the listed ROIs.
| It results a more expensive query, but its more powerful.
| It also enables you to require a minimum number of connections in the given
| ``inputRois`` or ``outputRois`` using the ``min_roi_inputs`` and ``min_roi_outputs``
| criteria.
|
| In either case, use use ``roi_req`` to specify whether a neuron must match just
| one (``any``) of the listed ROIs, or ``all`` of them.
|
| Args:
| matchvar (str):
| An arbitrary cypher variable name to use when this
| ``NeuronCriteria`` is used to construct cypher queries.
| To help catch errors (such as accidentally passing a ``type`` or
| ``instance`` name in the wrong argument position), we require that
| ``matchvar`` begin with a lowercase letter.
|
| bodyId (int or list of ints):
| List of bodyId values.
|
| instance (str or list of str):
| If ``regex=True``, then the instance will be matched as a regular expression.
| Otherwise, only exact matches are found. To search for neurons with no instance
| at all, use ``instance=[None]``. If both ``type`` and ``instance`` criteria are
| supplied, any neuron that matches EITHER criteria will match the overall criteria.
|
| type (str or list of str):
| If ``regex=True``, then the type will be matched as a regular expression.
| Otherwise, only exact matches are found. To search for neurons with no type
| at all, use ``type=[None]``. If both ``type`` and ``instance`` criteria are
| supplied, any neuron that matches EITHER criteria will match the overall criteria.
|
| regex (bool):
| If ``True``, the ``instance`` and ``type`` arguments will be interpreted as
| regular expressions, rather than exact match strings.
|
| cellBodyFiber (str or list of str):
| Matches for the neuron ``cellBodyFiber`` field. To search for neurons
| with no CBF at all, use ``cellBodyFiber=[None]``.
|
| status (str or list of str):
| Matches for the neuron ``status`` field. To search for neurons with no status
| at all, use ``status=[None]``.
|
| cropped (bool):
| If given, restrict results to neurons that are cropped or not.
|
| min_pre (int):
| Exclude neurons that don't have at least this many t-bars (outputs) overall,
| regardless of how many t-bars exist in any particular ROI.
|
| min_post (int):
| Exclude neurons that don't have at least this many PSDs (inputs) overall,
| regardless of how many PSDs exist in any particular ROI.
|
| rois (str or list of str):
| ROIs that merely intersect the neuron, without specifying whether
| they're intersected by input or output synapses.
| If not provided, will be auto-set from ``inputRois`` and ``outputRois``.
|
| inputRois (str or list of str):
| Only Neurons which have inputs in EVERY one of the given ROIs will be matched.
| ``regex`` does not apply to this parameter.
|
| outputRois (str or list of str):
| Only Neurons which have outputs in EVERY one of the given ROIs will be matched.
| ``regex`` does not apply to this parameter.
|
| min_roi_inputs (int):
| How many input (post) synapses a neuron must have in each ROI to satisfy the
| ``inputRois`` criteria. Can only be used if you provided ``inputRois``.
|
| min_roi_outputs (int):
| How many output (pre) synapses a neuron must have in each ROI to satisfy the
| ``outputRois`` criteria. Can only be used if you provided ``outputRois``.
|
| roi_req (Either ``'any'`` or ``'all'``):
| Whether a neuron must intersect all of the listed input/output ROIs, or any of the listed input/output ROIs.
| When using 'any', each neuron must still match at least one input AND at least one output ROI.
|
| label (Either ``'Neuron'`` or ``'Segment'``):
| Which node label to match with.
| (In neuprint, all ``Neuron`` nodes are also ``Segment`` nodes.)
| By default, ``'Neuron'`` is used, unless you provided a non-empty ``bodyId`` list.
| In that case, ``'Segment'`` is the default. (It's assumed you're really interested
| in the bodies you explicitly listed, whether or not they have the ``'Neuron'`` label.)
|
| client (:py:class:`neuprint.client.Client`):
| Used to validate ROI names.
| If not provided, the global default ``Client`` will be used.
|
| __repr__(self)
| Return repr(self).
|
| all_conditions(self, *vars, prefix=0, comments=True)
|
| basic_conditions(self, prefix=0, comments=True)
| Construct a WHERE clause based on the basic conditions
| in this criteria (i.e. everything except for the "directed ROI" conditions.)
|
| basic_exprs(self)
| Return the list of expressions that correspond
| to the members in this NeuronCriteria object.
| They're intended be combined (via 'AND') in
| the WHERE clause of a cypher query.
|
| bodyId_expr(self)
|
| cbf_expr(self)
|
| cropped_expr(self)
|
| directed_rois_condition(self, *vars, prefix=0, comments=True)
| Construct the ```WITH...WHERE``` statements that apply the "directed ROI"
| conditions specified by this criteria's ``inputRois`` and ``outputRois``
| members.
|
| These conditions are expensive to evaluate, so it's usually a good
| idea to position them LAST in your cypher query, once the result set
| has already been narrowed down by eariler filters.
|
| global_vars(self)
|
| global_with(self, *vars, prefix=0)
|
| instance_expr(self)
|
| post_expr(self)
|
| pre_expr(self)
|
| rois_expr(self)
|
| status_expr(self)
|
| type_expr(self)
|
| typeinst_expr(self)
| Unlike all other fields, type and instance OR'd together.
| Either match satisfies the criteria.
|
| ----------------------------------------------------------------------
| Class methods defined here:
|
| combined_conditions(neuron_conditions, vars=[], prefix=0, comments=True) from builtins.type
| Combine the conditions from multiple NeuronCriteria into a single string,
| putting the "cheap" conditions first and the "expensive" conditions last.
| (That is, basic conditions first and the directed ROI conditions last.)
|
| combined_global_with(neuron_conditions, vars=[], prefix=0) from builtins.type
|
| ----------------------------------------------------------------------
| Data descriptors defined here:
|
| __dict__
| dictionary for instance variables (if defined)
|
| __weakref__
| list of weak references to the object (if defined)
|
| ----------------------------------------------------------------------
| Data and other attributes defined here:
|
| MAX_LITERAL_LENGTH = 3
|
| __hash__ = None
Fetching neurons
Let’s say we want to find all antennnal lobe projection neurons (PNs). Their type nomenclature adherese to {glomerulus}_{lineage}PN (e.g. DA1_lPN)for uniglomerular PNs and a M_{lineage}PN{tract}{type} (e.g. M_vPNml50 = “multiglomerular ventral lineage PN mediolateral tract type 50) for multiglomerular PNs.
To get them all, we need to use regex patterns (see this cheatsheet):
# Define the filter criteria
nc = neu.NeuronCriteria(type='.*?_.*?PN.*?', regex=True)
# Get general info for these neurons
pns, roi_info = neu.fetch_neurons(nc)
print(f'{pns.shape[0]} PNs found.')
pns.head()
337 PNs found.
| bodyId | instance | type | pre | post | size | status | cropped | statusLabel | cellBodyFiber | somaRadius | somaLocation | inputRois | outputRois | roiInfo | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 294792184 | M_vPNml53_R | M_vPNml53 | 92 | 344 | 420662445 | Traced | False | Roughly traced | AVM04 | 336.5 | [18923, 34319, 35424] | [AL(R), AL-D(R), AL-DA2(R), AL-DA4m(R), AL-DC1... | [AL(R), AL-DC1(R), LH(R), PLP(R), SIP(R), SLP(... | {'SNP(R)': {'pre': 70, 'post': 155, 'downstrea... |
| 1 | 329599710 | M_lvPNm32_R | M_lvPNm32 | 247 | 285 | 343478957 | Traced | False | Roughly traced | AVM06 | NaN | None | [AL(R), AL-DC4(R), AL-DL2v(R), AL-DM1(R), AL-D... | [AL(R), AL-DL2v(R), AL-DM1(R), AL-DM4(R), AL-D... | {'SNP(R)': {'pre': 180, 'post': 93, 'downstrea... |
| 2 | 417199910 | M_lvPNm36_R | M_lvPNm36 | 162 | 347 | 387058559 | Traced | False | Roughly traced | AVM06 | 351.5 | [13823, 33925, 34176] | [AL(R), AL-DL5(R), AL-DM4(R), AL-DP1m(R), AL-V... | [AL(R), AL-DL5(R), AL-DM4(R), AL-VP1d(R), AL-V... | {'SNP(R)': {'pre': 156, 'post': 95, 'downstrea... |
| 3 | 480927537 | M_vPNml70_R | M_vPNml70 | 82 | 276 | 240153322 | Traced | False | Roughly traced | AVM04 | NaN | None | [AL(R), AL-DA2(R), AL-DA4l(R), AL-DA4m(R), AL-... | [LH(R), SLP(R), SNP(R)] | {'SNP(R)': {'pre': 15, 'post': 18, 'downstream... |
| 4 | 481268653 | M_vPNml89_R | M_vPNml89 | 146 | 58 | 265085609 | Traced | False | Roughly traced | AVM04 | NaN | None | [AL(R), AL-VC3l(R), AL-VC4(R), AL-VP1m(R), LH(... | [LH(R), SLP(R), SNP(R)] | {'SNP(R)': {'pre': 10, 'post': 2, 'downstream'... |
# Check that the regex did not have any accidental by-catch
pns['type'].unique()
array(['M_vPNml53', 'M_lvPNm32', 'M_lvPNm36', 'M_vPNml70', 'M_vPNml89',
'VP1l+_lvPN', 'M_vPNml69', 'DM1_lPN', 'DM4_vPN', 'M_vPNml79',
'VP4+_vPN', 'DA4l_adPN', 'M_vPNml87', 'DM4_adPN', 'M_vPNml83',
'VA5_lPN', 'DA4m_adPN', 'M_lvPNm24', 'M_vPNml85', 'VP1l+VP3_ilPN',
'M_vPNml77', 'M_vPNml84', 'VC1_lPN', 'M_lvPNm39', 'M_vPNml50',
'DM2_lPN', 'VC5_lvPN', 'M_vPNml88', 'M_vPNml58', 'VP4_vPN',
'DP1m_vPN', 'DP1m_adPN', 'DM5_lPN', 'VC5_adPN', 'M_vPNml80',
'M_lvPNm25', 'VC3m_lvPN', 'VP3+_vPN', 'VP1m+_lvPN', 'DA3_adPN',
'V_l2PN', 'M_vPNml56', 'VC3l_adPN', 'VM7v_adPN', 'DL5_adPN',
'VM4_adPN', 'VM2_adPN', 'M_lvPNm40', 'DC4_vPN', 'V_ilPN',
'M_vPNml74', 'Z_lvPNm1', 'DA1_lPN', 'DP1l_adPN', 'VM4_lvPN',
'M_vPNml71', 'DP1l_vPN', 'M_lvPNm41', 'M_spPN5t10', 'DA1_vPN',
'VC4_adPN', 'DM3_adPN', 'M_lvPNm45', 'VL1_vPN', 'M_lvPNm44',
'M_vPNml78', 'M_vPNml67', 'M_adPNm5', 'M_smPNm1', 'DM6_adPN',
'DL2d_adPN', 'M_adPNm6', 'M_adPNm8', 'M_lvPNm43', 'Z_vPNml1',
'M_vPNml59', 'DA2_lPN', 'M_lPNm11A', 'M_vPNml52', 'DL2d_vPN',
'VL2p_vPN', 'VA1d_adPN', 'M_lPNm11B', 'M_lvPNm48', 'M_lPNm11C',
'M_lvPNm42', 'VA1v_vPN', 'M_vPNml68', 'M_vPNml55', 'M_vPNml62',
'VL2a_vPN', 'M_vPNml60', 'M_vPNml65', 'VM5d_adPN', 'M_l2PNm16',
'M_vPNml61', 'M_vPNml57', 'M_vPNml64', 'M_lv2PN9t49',
'VP2+VC5_l2PN', 'M_spPN4t9', 'M_vPNml66', 'M_vPNml75', 'M_vPNml63',
'M_vPNml72', 'M_lvPNm38', 'D_adPN', 'M_vPNml76', 'M_vPNml54',
'DM3_vPN', 'M_vPNml86', 'DL3_lPN', 'VA4_lPN', 'VP1d_il2PN',
'DC1_adPN', 'M_l2PN3t18', 'M_lvPNm35', 'DL4_adPN', 'M_lvPNm28',
'M_lvPNm27', 'M_ilPNm90', 'M_l2PNl20', 'M_lvPNm29', 'VA7l_adPN',
'M_lPNm13', 'M_l2PNl21', 'DL1_adPN', 'M_imPNl92', 'M_vPNml73',
'M_ilPN8t91', 'M_l2PNm14', 'VP1d+VP4_l2PN1', 'M_lvPNm26',
'DL2v_adPN', 'VP3+VP1l_ivPN', 'M_lvPNm33', 'VA1v_adPN',
'VP3+_l2PN', 'M_l2PN10t19', 'VP4+VL1_l2PN', 'M_l2PNl22',
'M_l2PNm15', 'M_lPNm11D', 'MZ_lv2PN', 'DC2_adPN', 'M_lvPNm46',
'VC2_lPN', 'VM1_lPN', 'VM3_adPN', 'VM7d_adPN', 'M_lvPNm47',
'M_lPNm12', 'DC3_adPN', 'VP2+_adPN', 'VP1m+VP2_lvPN2',
'VP1m+VP2_lvPN1', 'VA6_adPN', 'VA7m_lPN', 'M_adPNm7', 'M_adPNm4',
'VA1d_vPN', 'VA3_adPN', 'VL1_ilPN', 'M_l2PNl23', 'M_lvPNm31',
'VP1m+VP5_ilPN', 'VL2p_adPN', 'MZ_lvPN', 'VP2_adPN', 'VA2_adPN',
'VM5v_adPN', 'VP5+VP2_l2PN', 'VP5+VP3_l2PN', 'VP5+_l2PN',
'M_vPNml51', 'M_smPN6t2', 'M_lvPNm37', 'M_vPNml82', 'M_adPNm3',
'VP1m_l2PN', 'DC4_adPN', 'VP5+Z_adPN', 'VL2a_adPN', 'VP2_l2PN',
'M_lvPNm34', 'VP2+Z_lvPN', 'M_lvPNm30', 'M_l2PNm17', 'M_vPNml81',
'VP1d+VP4_l2PN2'], dtype=object)
Fetching synaptic partners
Looks good! Next: What’s downstream of those PNs?
ds = neu.fetch_simple_connections(upstream_criteria=neu.NeuronCriteria(bodyId=pns.bodyId.values))
ds.head()
| bodyId_pre | bodyId_post | weight | type_pre | type_post | instance_pre | instance_post | conn_roiInfo | |
|---|---|---|---|---|---|---|---|---|
| 0 | 635062078 | 1671292719 | 390 | DP1m_adPN | lLN2T_c | DP1m_adPN_R | lLN2T_c(Tortuous)_R | {'AL(R)': {'pre': 390, 'post': 390}, 'AL-DP1m(... |
| 1 | 635062078 | 1704347707 | 326 | DP1m_adPN | lLN2T_c | DP1m_adPN_R | lLN2T_c(Tortuous)_R | {'AL(R)': {'pre': 324, 'post': 324}, 'AL-DP1m(... |
| 2 | 542634818 | 1704347707 | 322 | DM1_lPN | lLN2T_c | DM1_lPN_R | lLN2T_c(Tortuous)_R | {'AL(R)': {'pre': 322, 'post': 322}, 'AL-DM1(R... |
| 3 | 635062078 | 1640922516 | 320 | DP1m_adPN | lLN2T_e | DP1m_adPN_R | lLN2T_e(Tortuous)_R | {'AL(R)': {'pre': 317, 'post': 316}, 'AL-DP1m(... |
| 4 | 724816115 | 1670916819 | 318 | DP1l_adPN | lLN2P_a | DP1l_adPN_R | lLN2P_a(Patchy)_R | {'AL(R)': {'pre': 318, 'post': 318}, 'AL-DP1l(... |
Each row is now a connections from a single up- to a single downstream neuron. The “weight” is the number of synapses between the pre- and the postsynaptic neuron. Let’s simplify by grouping by type:
by_type = ds.groupby(['type_pre', 'type_post'], as_index=False).weight.sum()
by_type.sort_values('weight', ascending=False, inplace=True)
by_type.reset_index(drop=True, inplace=True)
by_type.head()
| type_pre | type_post | weight | |
|---|---|---|---|
| 0 | DC3_adPN | KCg-m | 3670 |
| 1 | VM5d_adPN | KCg-m | 3219 |
| 2 | DC1_adPN | KCg-m | 3215 |
| 3 | VL2a_adPN | KCg-m | 3096 |
| 4 | DA1_lPN | KCg-m | 3078 |
The strongest connections are between PNs and Kenyon Cells (KCs). That’s little surprising since there are thousands of KCs. For the sake of the argument let’s say we want to know where these connections occur:
adj, roi_info2 = neu.fetch_adjacencies(sources=neu.NeuronCriteria(bodyId=pns.bodyId.values),
targets=neu.NeuronCriteria(type='KC.*?', regex=True))
roi_info2.head()
0%| | 0/2 [00:00<?, ?it/s]
| bodyId_pre | bodyId_post | roi | weight | |
|---|---|---|---|---|
| 0 | 542634818 | 301314208 | CA(R) | 6 |
| 1 | 542634818 | 331999156 | CA(R) | 1 |
| 2 | 542634818 | 332344592 | CA(R) | 2 |
| 3 | 542634818 | 332344908 | CA(R) | 9 |
| 4 | 542634818 | 332353106 | CA(R) | 13 |
# Group by region of interest (ROI)
by_roi = roi_info2.groupby('roi').weight.sum()
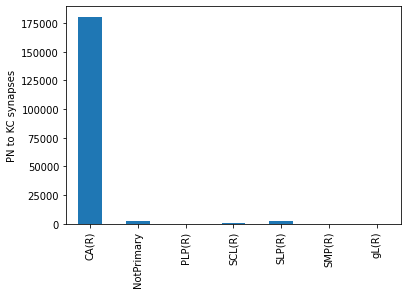
by_roi.head()
roi
CA(R) 180526
NotPrimary 2737
PLP(R) 11
SCL(R) 498
SLP(R) 2008
Name: weight, dtype: int64
ax = by_roi.plot.bar()
ax.set_xlabel('')
ax.set_ylabel('PN to KC synapses')
Text(0, 0.5, 'PN to KC synapses')
Querying paths
Let’s say we want to find out how to go from a PN (second order olfactory neurons) all the way to a descending neuron (presumably leading to motor neurons in the VNC).
# First fetch the DNs
dns, _ = neu.fetch_neurons(neu.NeuronCriteria(type='(.*DN[^1]{0,}.*|Giant Fiber)', regex=True))
dns.head()
| bodyId | instance | type | pre | post | size | status | cropped | statusLabel | cellBodyFiber | somaRadius | somaLocation | inputRois | outputRois | roiInfo | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 264083994 | DN1a_R | DN1a | 394 | 1231 | 1270566035 | Traced | False | Roughly traced | PDM10 | 270.0 | [11339, 22506, 4104] | [AME(R), CA(R), INP, MB(+ACA)(R), MB(R), OL(R)... | [AME(R), CA(R), INP, MB(+ACA)(R), MB(R), OL(R)... | {'SNP(R)': {'pre': 231, 'post': 998, 'downstre... |
| 1 | 295063181 | DNES2_R | DNES2 | 1 | 584 | 2051016758 | Traced | False | Roughly traced | PDM31 | 427.0 | [6063, 21133, 5000] | [CA(R), MB(+ACA)(R), MB(R), SLP(R), SMP(L), SM... | [SMP(R), SNP(R)] | {'SNP(R)': {'pre': 1, 'post': 561, 'downstream... |
| 2 | 324846570 | DN1pA_R | DN1pA | 184 | 445 | 800928414 | Traced | False | Roughly traced | PDM24 | 278.0 | [17791, 19036, 5000] | [SLP(R), SMP(L), SMP(R), SNP(L), SNP(R)] | [SLP(R), SMP(L), SMP(R), SNP(L), SNP(R)] | {'SNP(R)': {'pre': 97, 'post': 364, 'downstrea... |
| 3 | 325529237 | DN1pA_R | DN1pA | 201 | 436 | 790247619 | Traced | False | Roughly traced | PDM24 | 339.0 | [17387, 19226, 5776] | [SLP(R), SMP(L), SMP(R), SNP(L), SNP(R)] | [SLP(R), SMP(L), SMP(R), SNP(L), SNP(R)] | {'SNP(R)': {'pre': 116, 'post': 366, 'downstre... |
| 4 | 386834269 | DN1pB_R | DN1pB | 570 | 1050 | 1820640251 | Traced | False | Roughly traced | PDM24 | 357.0 | [18893, 20415, 3856] | [AOTU(R), INP, PLP(R), SCL(R), SIP(R), SLP(R),... | [AOTU(R), INP, PLP(R), SCL(R), SIP(R), SLP(R),... | {'SNP(R)': {'pre': 425, 'post': 856, 'downstre... |
Neuprint lets you query paths from a single source to a single target. For multi-source or -target queries, your best bet is to download the entire graph and run the queries locally using networkx or igraph.
# Find all paths from A PN to A DNs
paths = neu.fetch_shortest_paths(upstream_bodyId=pns.bodyId.values[0],
downstream_bodyId=dns.bodyId.values[0],
min_weight=10)
paths
| path | bodyId | type | weight | |
|---|---|---|---|---|
| 0 | 0 | 294792184 | M_vPNml53 | 0 |
| 1 | 0 | 5813057148 | SLP387 | 16 |
| 2 | 0 | 295478082 | SLP359 | 58 |
| 3 | 0 | 357224041 | LHPV5l1 | 21 |
| 4 | 0 | 388881226 | LHPV6m1 | 10 |
| 5 | 0 | 264083994 | DN1a | 18 |
| 6 | 1 | 294792184 | M_vPNml53 | 0 |
| 7 | 1 | 5813057148 | SLP387 | 16 |
| 8 | 1 | 295473947 | SLP359 | 65 |
| 9 | 1 | 357224041 | LHPV5l1 | 14 |
| 10 | 1 | 388881226 | LHPV6m1 | 10 |
| 11 | 1 | 264083994 | DN1a | 18 |
| 12 | 2 | 294792184 | M_vPNml53 | 0 |
| 13 | 2 | 5813057148 | SLP387 | 16 |
| 14 | 2 | 5813098375 | SLP347 | 10 |
| 15 | 2 | 5813071288 | SMP297 | 13 |
| 16 | 2 | 417558532 | SMP421 | 16 |
| 17 | 2 | 264083994 | DN1a | 13 |
| 18 | 3 | 294792184 | M_vPNml53 | 0 |
| 19 | 3 | 5813057148 | SLP387 | 16 |
| 20 | 3 | 296168382 | SLP347 | 21 |
| 21 | 3 | 5813071288 | SMP297 | 21 |
| 22 | 3 | 417558532 | SMP421 | 16 |
| 23 | 3 | 264083994 | DN1a | 13 |
So it looks like there are three separate 7-hop paths to go from M_vPNml53 to DN1a. Let’s plot a graph for this.
Plotting graphs
There are various ways of plotting static graphs. In theory Jupyter notebooks lend themselves to interactive graphs too but unfortunately DeepNote does not yet support the required libraries (e.g. ipywidgets). That being said: if you want to run this locally or on Google colab, check out ipycytoscape.
There are numerous options to do this but we will use networkx to plot a static graph:
import networkx as nx
import numpy as np
# Initialize the graph
G = nx.DiGraph()
# Generate edges from the paths
edges = []
for p in paths.path.unique():
this_path = paths.loc[(paths.path == p)]
this_edges = list(zip(this_path.values[:-1], this_path.values[1:]))
for i in range(this_path.shape[0] - 1):
edges.append([this_path.bodyId.values[i], this_path.bodyId.values[i + 1], this_path.weight.values[i + 1]])
# Add the edges
G.add_weighted_edges_from(edges)
# Add some names to the nodes
nx.set_node_attributes(G, paths.set_index('bodyId')['type'].to_dict(), name='name')
```python
```python
import matplotlib.pyplot as plt
# Draw using a simple force-directed layout
pos = nx.kamada_kawai_layout(G)
# We could draw everything in one step but this way we have more control over the plot
fig, ax = plt.subplots(figsize=(10, 10))
# Draw nodes
nx.draw_networkx_nodes(G, pos=pos, ax=ax)
# Draw edges
weights = np.array([e[2]['weight'] for e in G.edges(data=True)])
nx.draw_networkx_edges(G, pos=pos, width=(weights / 12).tolist())
# Add node labels
nx.draw_networkx_labels(G, pos=pos, labels=dict(G.nodes('name')), font_size=14)
# Turn axes of
ax.set_axis_off()
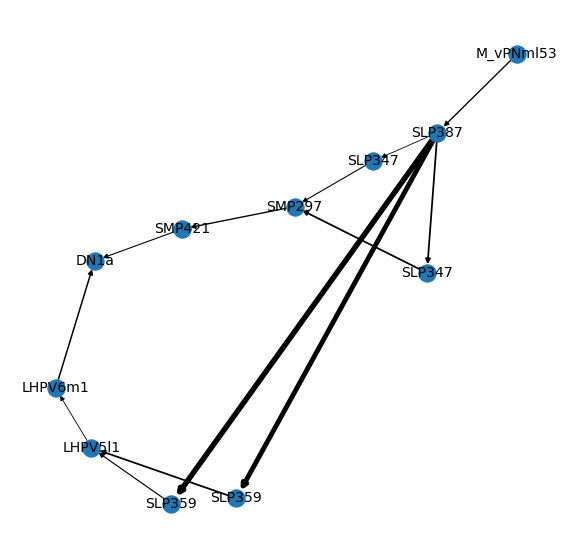
In general, I recommend exporting your graph to e.g. graphml and importing it into e.g. cytoscape if you want to explorate an interactive network graph.
nx.write_gml(G, "my_graph.gml")
<networkx.classes.digraph.DiGraph at 0x7f4ddb5c4d50>
G
<networkx.classes.digraph.DiGraph at 0x7f4ddb5c4d50>
Last but not least: let’s visualize the neurons involved!
Fetching meshes & skeletons
You can fetch skeletons as SWCs directly via neuprint-python. For visualization however it’s easiest to load neuron morphologies via navis. For that navis wraps neuprint-python and adds some convenience functions (see also the tutorial):
# Import the wrapped neuprint-python
# -> this exposes ALL base functions plus a couple navis-specific extras
import navis
import navis.interfaces.neuprint as neu
navis.set_pbars(jupyter=False)
client = neu.Client('https://neuprint.janelia.org', dataset='hemibrain:v1.1')
# Fetch neurons in the first path
nl = neu.fetch_skeletons(paths.loc[(paths.path == 0), 'bodyId'])
nl
| type | name | id | n_nodes | n_connectors | n_branches | n_leafs | cable_length | soma | units | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | navis.TreeNeuron | M_vPNml53_R | 294792184 | 3670 | 436 | 180 | 190 | 187780.664745 | 3649 | 8 nanometer |
| 1 | navis.TreeNeuron | DN1a_R | 264083994 | 7744 | 1625 | 813 | 841 | 349542.406630 | 7303 | 8 nanometer |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 4 | navis.TreeNeuron | SLP387_R | 5813057148 | 6776 | 1146 | 584 | 605 | 294831.692265 | 3 | 8 nanometer |
| 5 | navis.TreeNeuron | LHPV5l1_R | 357224041 | 19708 | 4748 | 1290 | 1336 | 856851.604331 | 7900 | 8 nanometer |
# Let's also get some ROI meshes
al = neu.fetch_roi('AL(R)')
lh = neu.fetch_roi('LH(R)')
ca = neu.fetch_roi('CA(R)')
# Plot
navis.plot3d([nl, lh, al, ca], width=1100)
Feedback
Was this page helpful?
Glad to hear it! Please tell us how we can improve.
Sorry to hear that. Please tell us how we can improve.