This is the multi-page printable view of this section. Click here to print.
About Virtual Fly Brain
- 1: How to cite us
- 2: Privacy Notice
- 3: What is Virtual Fly Brain?
- 4: Virtual Fly Brain Members and Their Contributions
- 5: Funding
- 6: Collaborators
- 7: Contributors
- 8: Publications
- 9: VFB citation bibtex
- 10: Contact us
- 11: Archived sites hosted by Virtual Fly Brain
- 11.1: ABD1.5 CATMAID Data Viewer
- 11.2: BrainTrap
- 11.3: FAFB CATMAID Data Viewer
- 11.4: FANC CATMAID Data Viewer
- 11.5: FLYBRAIN Neuron Database (NDB)
- 11.6: IAV-ROBO CATMAID Data Viewer
- 11.7: IAV-TNT CATMAID Data Viewer
- 11.8: L1EM CATMAID Data Viewer
1 - How to cite us
How to Cite Virtual Fly Brain
If you use Virtual Fly Brain data, tools, or resources in your research, please cite our work using the citation formats below. Proper citation helps support continued development and funding of this resource.
Primary Citation
Use this citation when referencing Virtual Fly Brain in general, including use of the website, data, or tools:
APA
Court, R., Costa, M., Pilgrim, C., Millburn, G., Holmes, A., McLachlan, A., Larkin, A., Matentzoglu, N., Kir, H., Parkinson, H., Brown, N. H., O’Kane, C. J., Armstrong, J. D., Jefferis, G. S. X. E., & Osumi-Sutherland, D. (2023). Virtual Fly Brain—An interactive atlas of the Drosophila nervous system. Frontiers in Physiology, 14. https://doi.org/10.3389/fphys.2023.1076533
MLA
Court, Robert, et al. “Virtual Fly Brain—An Interactive Atlas of the Drosophila Nervous System.” Frontiers in Physiology, vol. 14, 2023, https://doi.org/10.3389/fphys.2023.1076533.
Chicago
Court, Robert, Costa, Marta, Pilgrim, Clare, Millburn, Gillian, Holmes, Alex, McLachlan, Alex, Larkin, Aoife et al. “Virtual Fly Brain—An interactive atlas of the Drosophila nervous system.” Frontiers in Physiology 14, (2023). https://doi.org/10.3389/fphys.2023.1076533.
BibTeX
@article{10.3389/fphys.2023.1076533,
author = {Robert Court and Marta Costa and Clare Pilgrim and Gillian Millburn and Alex Holmes and Alex McLachlan and Aoife Larkin and Nicolas Matentzoglu and Huseyin Kir and Helen Parkinson and Nicolas H. Brown and Cahir J. O'Kane and J. Douglas Armstrong and Gregory S. X. E. Jefferis and David Osumi-Sutherland},
title = "{Virtual Fly Brain—An interactive atlas of the Drosophila nervous system}",
journal = {Frontiers in Physiology},
volume = {14},
publisher = {Frontiers Media {SA}},
year = {2023},
month = {jan},
abstract = "{As a model organism, Drosophila is uniquely placed to contribute to our understanding of how brains control complex behavior. Not only does it have complex adaptive behaviors, but also a uniquely powerful genetic toolkit, increasingly complete dense connectomic maps of the central nervous system and a rapidly growing set of transcriptomic profiles of cell types. But this also poses a challenge: Given the massive amounts of available data, how are researchers to Find, Access, Integrate and Reuse (FAIR) relevant data in order to develop an integrated anatomical and molecular picture of circuits, inform hypothesis generation, and find reagents for experiments to test these hypotheses? The Virtual Fly Brain (virtualflybrain.org) web application & API provide a solution to this problem, using FAIR principles to integrate 3D images of neurons and brain regions, connectomics, transcriptomics and reagent expression data covering the whole CNS in both larva and adult. Users can search for neurons, neuroanatomy and reagents by name, location, or connectivity, via text search, clicking on 3D images, search-by-image, and queries by type (e.g., dopaminergic neuron) or properties (e.g., synaptic input in the antennal lobe). Returned results include cross-registered 3D images that can be explored in linked 2D and 3D browsers or downloaded under open licenses, and extensive descriptions of cell types and regions curated from the literature. These solutions are potentially extensible to cover similar atlasing and data integration challenges in vertebrates.}",
issn = {1664-042X},
doi = {https://doi.org/10.3389/fphys.2023.1076533},
url = {https://doi.org/10.3389/fphys.2023.1076533},
}
Additional Citations
Drosophila Anatomy Ontology
If you specifically use or reference the Drosophila anatomy ontology developed as part of Virtual Fly Brain, please also cite:
- Osumi-Sutherland, D., Reeve, S., Mungall, C. J., Neuhaus, F., Ruttenberg, A., Jefferis, G. S. and Armstrong, J. D. (2012). A strategy for building neuroanatomy ontologies
Acknowledgements
When appropriate, please also acknowledge the specific data sources integrated into Virtual Fly Brain that you have used, such as JFRC, FlyBase, FlyCircuit, and datasets from individual research groups.
2 - Privacy Notice
Virtual Fly Brain Privacy Notice
Last updated: February 2026
The Virtual Fly Brain (VFB) project is committed to protecting your privacy and keeping you informed about how your personal information is used. This privacy notice explains how we collect, use, and protect your personal data when you visit our website.
What information do we collect?
We collect information about your visit to our website through Google Analytics, a web analytics service provided by Google, Inc. This includes:
- Your IP address (anonymized)
- Browser type and version
- Operating system
- Referring website
- Pages visited and time spent on each page
- Geographic location (country/city level only)
We do not collect any personally identifiable information such as names, email addresses, or contact details unless you voluntarily provide them (for example, through our contact forms or feedback mechanisms).
How do we use this information?
The information collected helps us:
- Understand how our website is used
- Improve the user experience and content
- Identify technical issues
- Analyze website traffic patterns
- Support our research by understanding user interests in neuroanatomical data
Cookies and Website Privacy
We use Google Analytics cookies to collect this information. These are small text files stored on your device. You can control cookies through your browser settings, and you can opt out of Google Analytics tracking by visiting Google Analytics Opt-out.
For more information about Google’s privacy practices, please see Google’s Privacy Policy.
Legal Basis for Processing
Our processing of your personal data is based on:
- Legitimate interests: To improve our website and research services
- Consent: Where you have provided it for specific purposes
Data Sharing and Disclosure
We do not sell, rent, or trade your personal information. Information collected by Google Analytics may be processed by Google on servers located in the United States. Google is committed to complying with the EU-U.S. Privacy Shield framework.
Data Retention
Analytics data is retained for up to 26 months, after which it is automatically deleted.
Your Rights
Under data protection law, you have rights including:
- Access: Request a copy of your personal data
- Rectification: Correct inaccurate data
- Erasure: Request deletion of your data
- Restriction: Limit how we process your data
- Objection: Object to processing based on legitimate interests
- Portability: Receive your data in a structured format
Contact Us
If you have questions about this privacy notice or wish to exercise your rights, please contact:
Data Protection Officer
University of Edinburgh
Old College
South Bridge
Edinburgh EH8 9YL
Email: dpo@ed.ac.uk
VFB Project Team
Email: data@virtualflybrain.org
Changes to this Privacy Notice
We may update this privacy notice from time to time. Any changes will be posted on this page with an updated revision date.
This privacy notice is supported by the University of Edinburgh’s wider privacy policies. For more information, visit the University Data Protection webpages.
3 - What is Virtual Fly Brain?
Welcome to Virtual Fly Brain (VFB) - an interactive tool for neurobiologists to explore the detailed neuroanatomy, neuron connectivity, and gene expression of Drosophila melanogaster. Our goal is to make it easier for researchers to find relevant anatomical information and reagents.
We integrate the neuroanatomical and expression data from the published literature, as well as image datasets onto the same brain template, making it possible to run cross searches, find similar neurons, and compare image data on our 3D Viewer.
4 - Virtual Fly Brain Members and Their Contributions
Virtual Fly Brain Team
Virtual Fly Brain is developed and maintained by an interdisciplinary team of researchers, developers, and curators across multiple UK institutions. Our collaborative approach brings together expertise in informatics, neuroscience, ontology development, and data curation.
3D Viewer, Online Tools, Server, and Website

Institute for Adaptive and Neural Computation, School of Informatics, University of Edinburgh
- Robert Court (Lead DevOps)
- Douglas Armstrong (Current Project PI)
- Nestor Milyaev (2009-2012)
Ontology Editing, FlyBase Support, and Data Annotation
Department of Genetics, University of Cambridge
- Cahir O’Kane (Current Project PI)
- Michael Ashburner (Original PI and Grant Holder)
- Simon Reeve (2009-2011)
- Nicole Staudt (2015-2016)
- Alex Holmes (2017-2019)
Department of Physiology, Development and Neuroscience
- Clare Pilgrim (Senior Ontology Editor/Curator)
- Alex McLachlan (Curator/UX Tester)
- Aoife Larkin (Curator)
- Gillian Millburn (Senior Curator)
- Nick Brown (Current Project PI)
Department of Zoology, University of Cambridge
- Marta Costa (Current Project Co-I)
Image Processing, Registration, and NBLAST Neuron Search

MRC Laboratory of Molecular Biology, Cambridge
- Greg Jefferis (Current Project PI)
Schema Development & Web Development

Wellcome Sanger Institute, Cambridge
- David Osumi-Sutherland (Current Project Co-I)
European Bioinformatics Institute (EMBL-EBI), Cambridge
- Huseyin Kir (Developer)
- Nico Matentzoglu (Developer)
- Helen Parkinson (Current Project PI)
Software Engineering, Geppetto Customisation, Graphics Design

MetaCell
MetaCell collaborates with Virtual Fly Brain on the development and enhancement of the open source Geppetto visualization platform that powers our 3D brain viewer.
For questions about team members or collaboration opportunities, please contact us.
5 - Funding
Current Funding (2022-2027)
Wellcome Trust Grant 223741/Z/21/Z
- Duration: April 1, 2022 to March 31, 2026 (extended to March 31, 2027)
- Supporting ongoing VirtualFlyBrain development and maintenance
Historical Funding
Wellcome Trust
Grant 208379/Z/17/Z
- Duration: October 1, 2017 to October 1, 2021
“Virtual Fly Brain: a global informatics hub for Drosophila neurobiology”
- Grant 105023/D/14/Z: October 1, 2014 to January 31, 2018
- Grant 105023/A/14/Z: October 1, 2014 to January 31, 2018
- Grant 105023/C/14/Z: October 1, 2014 to September 30, 2017
- Grant 105023/B/14/Z: October 1, 2014 to September 30, 2017
UK Research Councils
Biotechnology and Biological Sciences Research Council (BBSRC)
- Grant BB/G02233X/1 (2009): “Standardising the representation of Drosophila anatomy and development for databases”
- Recipients: J. Douglas Armstrong, Michael Ashburner, Cahir O’Kane, David Osumi-Sutherland
Engineering and Physical Sciences Research Council (EPSRC)
- UK e-Science Theme Award supporting initial project establishment
- Recipient: J. Douglas Armstrong
Medical Research Council (MRC)
- Ongoing institutional support for technical infrastructure development
Other Sources
Isaac Newton Trust (University of Cambridge)
- Grant supporting David Osumi-Sutherland’s database work (2007)
- Grant supporting Marta Costa’s research: “Neuroinformatic identification of new types of neuron in the Drosophila brain” (October 2012 - September 2013)
Wellcome Trust - Cambridge Protein Trap Project
- Supporting BrainTrap database development
- Recipients: Kathryn Lilley, Steve Russell, Daniel St. Johnson
Institutional Partners
VirtualFlyBrain is a collaborative project between:
- University of Edinburgh - School of Informatics
- University of Cambridge - Multiple departments including Genetics, Physiology Development & Neuroscience, and Zoology
- EMBL’s European Bioinformatics Institute (EMBL-EBI) - Hinxton, UK
- Wellcome Sanger Institute - Hinxton, UK
- MRC Laboratory for Molecular Biology - Cambridge
Data Partnerships
VirtualFlyBrain integrates data from multiple sources through collaborative partnerships rather than direct funding arrangements:
- FlyBase Consortium
- FlyCircuit Project (National Tsing Hua University, Taiwan)
- Janelia Research Campus (HHMI, USA)
- Multiple international research laboratories
Site Visualization Tools
VirtualFlyBrain utilizes multiple visualization technologies:
Geppetto Framework
- Open source 3D visualization platform co-developed with MetaCell Corporation
- VirtualFlyBrain and MetaCell collaborate on ongoing development and improvements to the Geppetto ecosystem
Woolz Image Processing and IIP3D Server
- Developed by the MRC Human Genetics Unit, University of Edinburgh
- Provides high-performance image serving and client-side visualization tools
6 - Collaborators
Collaborators
The IIP3D server, Woolz software and client-side tools are developed by*
MRC Human Genetics Unit (MRC HGU): Richard Baldock, Nick Burton, Bill Hill, Zsolt Husz
(*) An on-going development of the client-side tools is done in collaboration between the MRC HGU and Edinburgh University
Visit the EMAGE gene expression database to see other tools the MRC HGU have developed.
Expression data is collaboratively curated by VFB and FlyBase. Phenotype data is curated by FlyBase. Expression and phenotype data displayed on VFB is stored and maintained by FlyBase.
7 - Contributors
Contributors
We would like to thank the following contributors for their help with this project:
8 - Publications
Publications
For more information on the technology behind the VFB website:
- Robert Court, Marta Costa, Clare Pilgrim, Gillian Millburn, Alex Holmes, Alex McLachlan, Aoife Larkin, Nicolas Matentzoglu, Huseyin Kir, Helen Parkinson, Nicolas H. Brown, Cahir J. O’Kane, J. Douglas Armstrong, Gregory S. X. E. Jefferis and David Osumi-Sutherland (2023). Virtual Fly Brain - An interactive atlas of the Drosophila nervous system. Frontiers in Physiology 14.
- Matteo Cantarelli, Boris Marin, Adrian Quintana, Matt Earnshaw, Robert Court, Padraig Gleeson, Salvador Dura-Bernal, R. Angus Silver, Giovanni Idili (2018). Geppetto: a reusable modular open platform for exploring neuroscience data and models. Philosophical Transactions of the Royal Society B: Biological Sciences 373.
- Hilmar Lapp, James P. Balhoff, Todd J. Vision (2017). Owlery: A flexible approach for the serving of OWL ontologies. bioRxiv.
- Milyaev, N., Osumi-Sutherland, D., Reeve, S., Burton, N., Baldock, R. A. and Armstrong, J. D. (2012). The Virtual Fly Brain browser and query interface. Bioinformatics 28, 411-5.
- Husz ZL, Burton N, Hill B, Milyaev N, Baldock RA:Web tools for large-scale 3D biological images and atlases. BMC Bioinformatics 13:122, 2012.
For details on the anatomy ontology:
- Osumi-Sutherland, D., Reeve, S., Mungall, C. J., Neuhaus, F., Ruttenberg, A., Jefferis, G. S. and Armstrong, J. D. (2012). A strategy for building neuroanatomy ontologies. Bioinformatics 28, 1262-1269.
9 - VFB citation bibtex
@article{Court_2023,
doi = {10.3389/fphys.2023.1076533},
url = {https://doi.org/10.3389/fphys.2023.1076533},
year = 2023,
month = {jan},
publisher = {Frontiers Media {SA}},
volume = {14},
author = {Robert Court and Marta Costa and Clare Pilgrim and Gillian Millburn and Alex Holmes and Alex McLachlan and Aoife Larkin and Nicolas Matentzoglu and Huseyin Kir and Helen Parkinson and Nicolas H. Brown and Cahir J. O'Kane and J. Douglas Armstrong and Gregory S. X. E. Jefferis and David Osumi-Sutherland},
title = {Virtual Fly Brain{\textemdash}An interactive atlas of the Drosophila nervous system}, journal = {Frontiers in Physiology}}
10 - Contact us
Contact Virtual Fly Brain
We welcome feedback, questions, and collaboration opportunities from the Drosophila research community. Please choose the most appropriate contact method for your inquiry below.
Technical Support & General Questions
For help with using Virtual Fly Brain, comments, suggestions, or general questions:
Public Support Forum: support@virtualflybrain.org
Note: This email goes to our public support forum, where questions and responses are visible to the community, helping other users with similar issues.
Before emailing: Please check our support forum archives to see if your question has already been answered.
Bug Reports & Feature Requests
To report technical issues, bugs, or request new features:
GitHub Issues: Report an issue
This allows our development team to track and address technical problems efficiently.
Data Contributions & Private Inquiries
For data inclusion discussions, collaboration proposals, or confidential matters that require direct communication with our team:
Private Email: data@virtualflybrain.org
Learn more about data contributions: Contribution Guidelines
Collaboration & Partnerships
Interested in collaborating with the Virtual Fly Brain team or discussing partnership opportunities?
Contact: data@virtualflybrain.org
Questions about Data Inclusion
For questions about adding your data:
Contact: data@virtualflybrain.org
Stay Connected
- BlueSky: Follow us for updates and announcements
- LinkedIn: Follow us for updates and announcements
- GitHub: VirtualFlyBrain organization - contribute to our open source development
11 - Archived sites hosted by Virtual Fly Brain
11.1 - ABD1.5 CATMAID Data Viewer
Access
The ABD1.5 CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance is accessible at: https://abd1.5.catmaid.virtualflybrain.org/
This resource provides wild-type reference data for the abdominal nerve cord of first instar Drosophila larvae, serving as a baseline for comparative connectomics studies. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publications
This dataset contains neuron reconstructions from multiple foundational connectomics studies:
-
Ohyama T, Schneider-Mizell CM, Fetter RD, Aleman JV, Franconville R, Rivera-Alba M, et al. (2015). A multilevel multimodal circuit enhances action selection in Drosophila. Nature, 520(7545), 633-639. https://doi.org/10.1038/nature14297
-
Schneider-Mizell CM, Gerhard S, Longair M, Kazimiers T, Li F, Zwart MF, et al. (2016). Quantitative neuroanatomy for connectomics in Drosophila. eLife, 5, e12059. https://doi.org/10.7554/eLife.12059
-
Valdes-Aleman J, Fetter RD, Sales EC, Heckman EL, Venkatasubramanian L, Doe CQ, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
Dataset Contents
The viewer provides access to:
- Serial section electron microscopy data of Drosophila 1st instar larval CNS (1.5-segment abdominal fraction, A2-A3 segments)
- Image resolution: 3.8 nm x 3.8 nm x 40 nm (x, y, z)
- Wild-type control neural tissue
- Hundreds of manually traced and reconstructed neurons
- Synaptic connectivity information
- Associated metadata and annotations
Reconstructed Neuron Types
This dataset includes extensive reconstructions of:
Sensory neurons:
- Mechanosensory chordotonal neurons (lch5 subtypes)
- Nociceptive multidendritic neurons (ddaC, ddaD, ddaE, vdaB, v’ada)
- Polymodal sensory neurons (v’ch, vchA/B, vchB/A)
Interneurons:
- Basin (Bs) interneurons (excitatory modulation)
- Proprioceptive and intersegmental interneurons
- Premotor interneurons (extensive IS premotor reconstructions)
- Motor circuit interneurons (Looper, Pseudolooper, Scepter, Bowl, Chair classes)
Motor neurons:
- Identified motor neurons (aCC, RP2)
- Putative motor neurons and muscle-innervating neurons
Other circuit elements:
- Intersegmental connections
- Spillover neurons (anterior and posterior processes)
- Additional circuit fragments and auxiliary neurons
Technical Details
The EM volume was prepared and imaged following protocols detailed in Ohyama et al. (2015). Neurons were reconstructed using CATMAID to obtain skeletonized structures and connectivity information. All synaptic connections represent chemical synapses.
Key neuron types identifiable in this dataset include:
- Mechanosensory chordotonal neurons (input sensory neurons)
- Basin (Bs) interneurons (excitatory integration)
- Drunken (Dr), Griddle (Gr), and Ladder (Ld) interneurons (inhibitory modulation)
- Nociceptive multidendritic neurons (noxious stimulus detection)
- Local and ascending interneurons
These neurons are uniquely identifiable based on:
- Nerve entry point of the main neurite into the neuropil
- Growth pattern of main axonal and dendritic branches
- Bilateral vs ipsilateral projections
- Position of terminal projections within the neuropil
Features
This CATMAID instance enables:
- Browser-based visualization of serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://abd1.5.catmaid.virtualflybrain.org/apis/
Citation Guidelines
When using this data, please cite the relevant research:
-
For mechanosensory and motor circuit data: Valdes-Aleman J, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
-
For foundational circuit architecture and quantitative neuroanatomy: Ohyama T, et al. (2015). A multilevel multimodal circuit enhances action selection in Drosophila. Nature, 520(7545), 633-639. https://doi.org/10.1038/nature14297
-
For quantitative neuroanatomy methodology: Schneider-Mizell CM, et al. (2016). Quantitative neuroanatomy for connectomics in Drosophila. eLife, 5, e12059. https://doi.org/10.7554/eLife.12059
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources.
11.2 - BrainTrap
https://braintrap.virtualflybrain.org/
Overview
BrainTrap was developed as part of a large-scale collaborative research project to collect protein expression location information from protein-trap lines generated by the Cambridge Protein Trap (CPT) project. The database provides detailed 3D confocal datasets showing protein expression patterns throughout the adult Drosophila brain.
Features
- Interactive web-based viewer for exploring 3D confocal datasets
- Full-size images throughout the brain volume can be viewed interactively
- Secondary immunohistochemical label (anti-brp) aids navigation and helps identify brain structures
- Searchable annotations linked to the FlyBase Drosophila anatomy ontology
- Anatomical search criteria can be specified using:
- Automatic term completion
- Hierarchical browser for the ontology
- Annotation provenance tracking with highlighted expression locations
- Downloadable confocal dataset files in original format
- XML format annotation data export
- Gene comparison tables for selected genes
Original Publication
Knowles-Barley S, Longair M and Armstrong JD (2010). BrainTrap: a database of 3D protein expression patterns in the Drosophila brain. Database, Vol. 2010, Article ID baq005, doi:10.1093/database/baq005 (https://doi.org/10.1093/database/baq005)
Funding
Original development was funded by:
- Engineering and Physical Sciences Research Council (EPSRC)
- Medical Research Council
- Biotechnology and Biological Sciences Research Council (BBSRC)
- British Society for Developmental Biology
- Society for Experimental Biology
- Virtual Fly Brain e-Science Institute (ESI) Theme
The fly lines were generated by the Cambridge Protein Trap Project, supported by a Wellcome Trust grant to Kathryn Lilley, Steve Russell and Daniel St. Johnson.
Current Maintenance
This resource is currently archived and maintained by Virtual Fly Brain (VFB).
Current website: https://braintrap.virtualflybrain.org/
Source code repository: https://github.com/VirtualFlyBrain/BrainTrap
Original Development
School of Informatics
The University of Edinburgh
Edinburgh, EH8 9AB, UK
Citation
If you use BrainTrap in your research, please cite:
Knowles-Barley S, Longair M and Armstrong JD (2010). BrainTrap: a database of 3D protein expression patterns in the Drosophila brain. Database, Vol. 2010, Article ID baq005, doi:10.1093/database/baq005 (https://doi.org/10.1093/database/baq005)
11.3 - FAFB CATMAID Data Viewer
Access
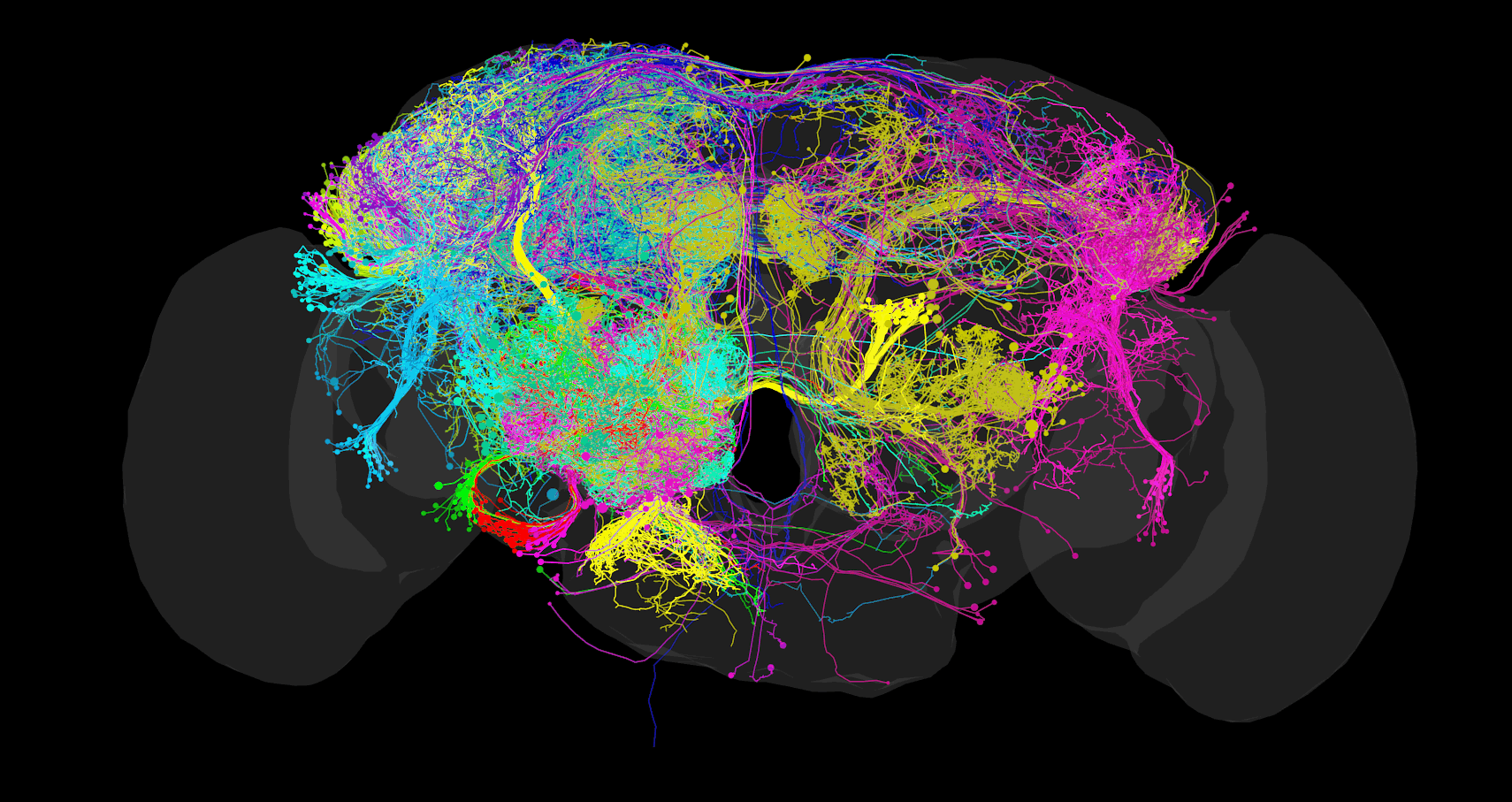
The FAFB CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance is accessible at: https://fafb.catmaid.virtualflybrain.org/
This resource provides access to the FAFB dataset and its associated neural reconstructions. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publications
Primary Resources
- FAFB Project: https://flyconnecto.me/
- EM Dataset: Zheng Z, et al. (2018) A Complete Electron Microscopy Volume of the Brain of Adult Drosophila melanogaster. Cell, 174(3), 730-743.e22. https://doi.org/10.1016/j.cell.2018.06.019
Contributing Publications
The database includes neurons traced and published in numerous studies. Each neuron is annotated with its source publication. Major contributing publications include:
- Circuit reorganization in the Drosophila mushroom body calyx accompanies memory consolidation (Baltruschat et al. 2021)
- Complete connectome of the Drosophila mushroom body provides insights into memory organization (Bates and Schlegel et al. 2020)
- The Connectome of the Adult Drosophila Mushroom Body Provides Insights into Function (Coates et al. 2020)
- Integration of the olfactory code across dendritic claws of single mushroom body neurons (Dolan and Belliart-Guérin et al. 2018)
- Neurogenetic dissection of the Drosophila lateral horn reveals major outputs, diverse behavioural functions, and interactions with the mushroom body (Dolan et al. 2019)
- Drosophila gustatory projections are segregated by taste modality and connectivity (Engert et al. 2022)
- Re-evaluation of learned information in Drosophila (Felsenberg et al. 2018)
- Motor neurons generate pose-targeted movements via proprioceptive sculpting (Gorko et al. 2024)
- Distinct subpopulations of mechanosensory chordotonal organ neurons elicit grooming of the fruit fly antennae (Hampel et al. 2020)
- A neural circuit for wind-guided olfactory navigation (Kim et al. 2020)
- Parallel encoding of sensory history and behavioral preference during Caenorhabditis elegans olfactory learning (Kind et al. 2021)
- A neural circuit for diurnal temperature preference in Drosophila (Marin et al. 2020)
- Spatial readout of visual looming in the central brain of Drosophila (Morimoto et al. 2020)
- Input Connectivity Reveals Additional Heterogeneity of Dopaminergic Reinforcement in Drosophila (Otto et al. 2020)
- A Neural Circuit Architecture for Angular Integration in Drosophila (Sayin et al. 2019)
- Taste quality and hunger interactions in a feeding sensorimotor circuit (Shiu et al. 2022)
- Generating parallel representations of position and identity in the olfactory system (Taisz et al. 2023)
- A synaptic threshold mechanism for computing escape decisions (Turner-Evans et al. 2020)
- Neural circuitry linking mating and egg laying in Drosophila females (Wang et al. 2020a)
- Circuit and Behavioral Mechanisms of Sexual Rejection by Drosophila Females (Wang et al. 2020b)
- Neural circuit mechanisms of sexual receptivity in Drosophila females (Wang et al. 2020c)
- Zhao et al. (2023)
- Structured sampling of olfactory input by the fly mushroom body (Zheng et al. 2022)
Full references for specific neurons can be found on Virtual Fly Brain by searching for their skeleton ID (skid).
Dataset Contents
The viewer provides access to:
- Complete electron microscopy volume of an adult Drosophila melanogaster brain
- Manually traced neuron reconstructions from multiple research groups
- Synaptic connectivity information
- Associated metadata and annotations
Features
This CATMAID instance enables:
- Browser-based visualization of the serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface with paper-based filtering
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://fafb.catmaid.virtualflybrain.org/apis/
Citation Guidelines
When using this data, please cite:
-
The FAFB EM dataset: Zheng Z, et al. (2018) A Complete Electron Microscopy Volume of the Brain of Adult Drosophila melanogaster. Cell, 174(3), 730-743.e22. https://doi.org/10.1016/j.cell.2018.06.019
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
-
The specific publication(s) associated with any neurons you analyze or reference (see Contributing Publications section)
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources.
11.4 - FANC CATMAID Data Viewer
Access
The FANC CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance provides two views of the data:
-
Original EM stack and tracings (Project ID 1): https://fanc.catmaid.virtualflybrain.org/
-
Neurons aligned to JRC2018 VNC female template (Project ID 2): https://fanc.catmaid.virtualflybrain.org/
This resource provides access to both the original FANC dataset and template-aligned reconstructions. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publication
The data is from the research published in:
Fushiki A, et al. (2021) A circuit mechanism for the propagation of waves of muscle contraction in Drosophila. Cell, 184(3), 759-774.e20. https://doi.org/10.1016/j.cell.2020.12.013
see: https://www.lee.hms.harvard.edu/resources
Dataset Contents
The viewer provides access to:
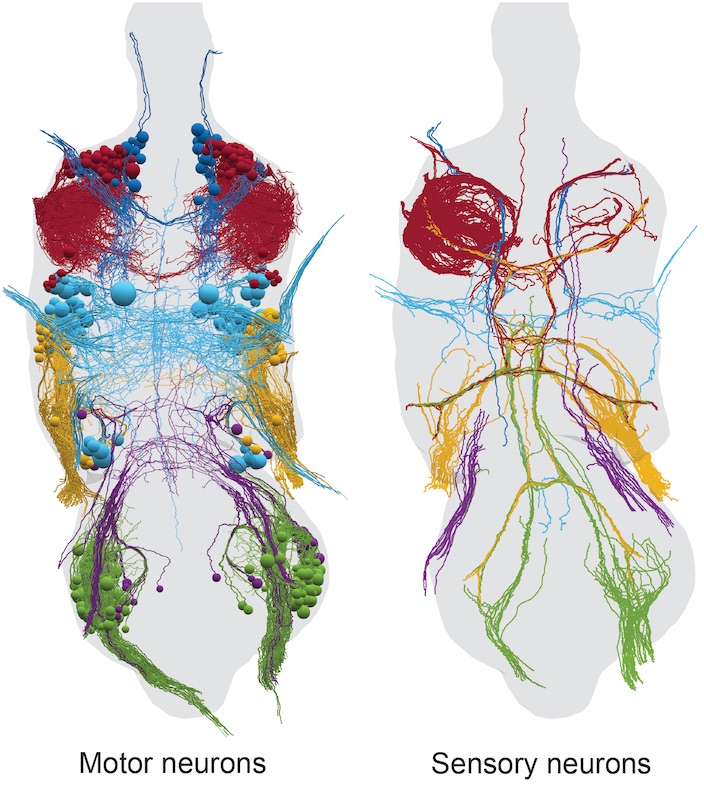
- Complete electron microscopy volume of an adult female Drosophila ventral nerve cord
- Manually traced neuron reconstructions in original EM space
- Template-aligned neurons in JRC2018 VNC female template space
- Synaptic connectivity information
- Associated metadata and annotations
Features
This CATMAID instance enables:
- Browser-based visualization of the serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions in both original and template space
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://fanc.catmaid.virtualflybrain.org/apis/
Note: Both views (original and aligned) can be accessed through the same API using different project IDs (pid).
Citation Guidelines
When using this data, please cite:
-
The FANC dataset: Fushiki A, et al. (2021) A circuit mechanism for the propagation of waves of muscle contraction in Drosophila. Cell, 184(3), 759-774.e20. https://doi.org/10.1016/j.cell.2020.12.013
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources. The aligned neurons are also available through the Virtual Fly Brain website.
11.5 - FLYBRAIN Neuron Database (NDB)
https://flybrain-ndb.virtualflybrain.org/
Database Structure
FLYBRAIN NDB is a relational database searchable by two main criteria:
1. Neuron Search
Search neurons by:
- Name (full or partial)
- Areas of arborizations
- Distribution of pre- or postsynaptic sites (where known)
- Keywords in documentation
2. Strain/Antibody Search
Search molecular markers including:
- Antibodies
- Drosophila strains (e.g., GAL4 enhancer-trap lines)
- Specific neural pathways
Data Categories
- Visual Neurons
- Olfactory Neurons
- Auditory Neurons
- Gustatory Neurons
- Central Complex Neurons
- Clonal Units
- Brain Region Definitions
- GAL4 and LexAV Strain Collections
Brain Explorer Features
The online brain browsing system allows users to:
- Rotate volume-rendered images
- Create sections
- Adjust magnification
- Browse confocal data
Terms of Use
Access
- FLYBRAIN NDB is freely available
- No login or registration required
Copyright
- All contents © 2008 FLYBRAIN NDB or their original publication
- Copyright is noted in respective images
Usage Rights
- Free use for non-profit, educational, and scientific purposes with proper citation
- Republication or commercial use requires consent from the FLYBRAIN NDB manager
Citation Guidelines
Citing the Database
When citing the database as a whole, use: “Flybrain Neuron Database (https://flybrain-ndb.VirtualFlyBrain.org)”
Citing Specific Records
Format: [Neuron type], [Author(s)], ([Year]) Flybrain Neuron Database (https://flybrain-ndb.VirtualFlyBrain.org), acc. [number]
Example: “LC10 neuron, Otsuna, H., and Ito, K. (2007) Flybrain Neuron Database (https://flybrain-ndb.VirtualFlyBrain.org), acc. 10010”
Important Notes:
- Include accession numbers when citing specific neurons
- Cite original publications where applicable
- Multiple contributors may exist for each record
Current Maintenance
This resource is currently archived and maintained by Virtual Fly Brain (VFB).
Source code repository: https://github.com/VirtualFlyBrain/flybrain-ndb
Original Development
© 2010 FLYBRAIN Neuron Database
Original Contact: Kazunori Shinomiya
11.6 - IAV-ROBO CATMAID Data Viewer
Access
The IAV-ROBO CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance is accessible at: https://iav-robo.catmaid.virtualflybrain.org/
This resource provides a direct view into the dataset used to study the organization of proprioceptive circuits in Drosophila larvae. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publication
This data is from the research published in:
Valdes-Aleman J, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
Dataset Contents
The viewer provides access to:
- Serial section electron microscopy data of Drosophila 1st instar larval CNS (1.5-segment fraction, A1/A2 segment)
- Image resolution: 3.8 nm x 3.8 nm x 40 nm (x, y, z)
- Genotype: w;; iav-GAL4/UAS-FraRobo
- Manually traced neuron reconstructions showing shifted mechanosensory projections
- Synaptic connectivity information
- Associated metadata and annotations
Technical Details
The EM volume was prepared and imaged following protocols detailed in Ohyama et al. (2015). Neurons were reconstructed using CATMAID to obtain skeletonized structures and connectivity information. All synaptic connections represent chemical synapses.
Despite the FraRobo-induced lateral shift in terminal projections, neurons remain uniquely identifiable based on:
- Nerve entry point of the main neurite into the neuropil
- Growth pattern of main axonal and dendritic branches
- Bilateral vs ipsilateral projections
For example, mechanosensory chordotonal neurons can be identified by their:
- Eight stereotypic entry points per hemisegment
- Most lateral position among sensory neurons in their nerve bundle
- Further lateral shift due to FraRobo expression
Features
This CATMAID instance enables:
- Browser-based visualization of the serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://iav-robo.catmaid.virtualflybrain.org/apis/
Citation Guidelines
When using this data, please cite both:
-
The original research: Valdes-Aleman J, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources.
11.7 - IAV-TNT CATMAID Data Viewer
Access
The IAV-TNT CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance is accessible at: https://iav-tnt.catmaid.virtualflybrain.org/
This resource provides a direct view into the dataset used to study the organization of proprioceptive circuits in Drosophila larvae. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publication
This data is from the research published in:
Valdes-Aleman J, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
Dataset Contents
The viewer provides access to:
- Serial section electron microscopy data of Drosophila 1st instar larval CNS (Control 1: whole-CNS volume, A1 segment)
- Image resolution: 3.8 nm x 3.8 nm x 40 nm (x, y, z)
- Manually traced neuron reconstructions
- Synaptic connectivity information
- Associated metadata and annotations
Technical Details
The EM volume was prepared and imaged following protocols detailed in Ohyama et al. (2015). Neurons were reconstructed using CATMAID to obtain skeletonized structures and connectivity information. All synaptic connections represent chemical synapses.
The neurons are uniquely identifiable based on key morphological features:
- Nerve entry point of the main neurite into the neuropil
- Growth pattern of main axonal and dendritic branches
- Bilateral vs ipsilateral projections
- Position of terminal projections within the medio-lateral, dorso-ventral and antero-posterior axes
Features
This CATMAID instance enables:
- Browser-based visualization of the serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://iav-tnt.catmaid.virtualflybrain.org/apis/
Citation Guidelines
When using this data, please cite both:
-
The original research: Valdes-Aleman J, et al. (2021). Comparative Connectomics Reveals How Partner Identity, Location, and Activity Specify Synaptic Connectivity in the Drosophila Motor System. Neuron, 109(1), 105-120.e7. https://doi.org/10.1016/j.neuron.2020.10.004
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources.
11.8 - L1EM CATMAID Data Viewer
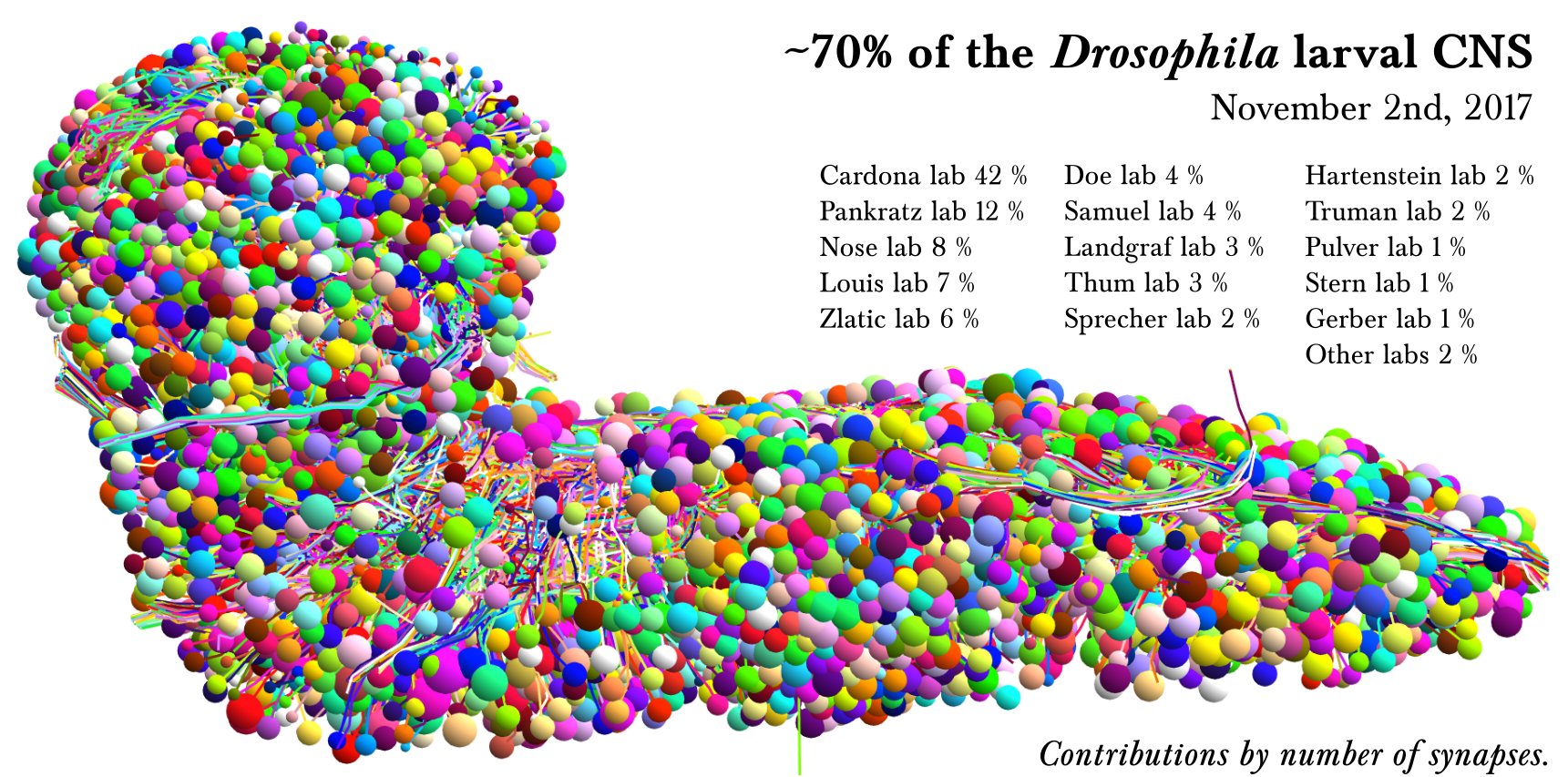

Access
The L1EM CATMAID instance is hosted and maintained by Virtual Fly Brain (VFB) as part of their mission to integrate and preserve key Drosophila neuroscience datasets. The instance is accessible at: https://l1em.catmaid.virtualflybrain.org/
This resource provides access to the L1EM dataset and its associated neural reconstructions. Virtual Fly Brain ensures its long-term availability to the research community.
Source Publication
Primary Resource
- L1EM Dataset: Winding M, et al. (2023) The connectome of an insect brain. Science, 379(6636):eadd9330. https://doi.org/10.1126/science.add9330
Contributing Publications
The database includes neurons traced and published in numerous studies. Each neuron is annotated with its source publication. Major contributing publications include:
- Andrade et al. (2019)
- Barnes et al. (2022)
- Berck, Khandelwal et al. (2016)
- Burgos et al. (2018)
- Carreira-Rosario, Arzan Zarin, Clark et al. (2018)
- Eichler, Li, Litwin-Kumar et al. (2017)
- Eschbach, Fushiki et al. (2020, 2020b)
- Fushiki et al. (2016)
- Gerhard et al. (2017)
- Heckscher et al. (2015)
- Hueckesfeld et al. (2020)
- Imambocus et al.
- Jovanic et al. (2019)
- Jovanic, Schneider-Mizell et al. (2016)
- Larderet, Fritsch et al. (2017)
- Mark et al. (2019)
- Miroschnikow et al. (2018)
- Ohyama, Schneider-Mizell et al. (2015)
- Schlegel et al. (2016)
- Takagi et al. (2017)
- Tastekin et al. (2018)
- Valdes-Aleman et al. (2021)
- Winding, Pedigo et al. (2023)
- Zarin, Mark et al. (2019)
- Zwart et al. (2016)
Full references for specific neurons can be found on Virtual Fly Brain by searching for their skeleton ID (skid).
Dataset Contents
The viewer provides access to:
- Complete electron microscopy volume of a Drosophila first instar larva brain
- Manually traced neuron reconstructions from multiple research groups
- Synaptic connectivity information
- Associated metadata and annotations
Features
This CATMAID instance enables:
- Browser-based visualization of the serial section electron microscopy data
- Navigation through image stacks
- Viewing of neuron reconstructions
- Analysis of synaptic connectivity
- Data export functionality
Available Tools
- Tracing tool for examining reconstructions
- Neuron search interface with paper-based filtering
- Connectivity analysis tools
- Skeleton visualization options
- API access for programmatic data retrieval (documentation: https://catmaid.readthedocs.io/en/stable/api.html): https://l1em.catmaid.virtualflybrain.org/apis/
Citation Guidelines
When using this data, please cite:
-
The L1EM dataset: Winding M, et al. (2023) The connectome of an insect brain. Science, 379(6636):eadd9330. https://doi.org/10.1126/science.add9330
-
The CATMAID platform: Saalfeld S, Cardona A, Hartenstein V, Tomančák P (2009) CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25(15): 1984-1986. https://doi.org/10.1093/bioinformatics/btp266
-
The specific publication(s) associated with any neurons you analyze or reference (see Contributing Publications section)
Maintenance & Support
This resource is archived, hosted, and maintained by Virtual Fly Brain (VFB - https://virtualflybrain.org) as part of their commitment to preserving and making accessible critical Drosophila neuroscience data resources.

